Cite as: Leliaert F, Smith DR, Moreau H, Herron MD, Verbruggen H, Delwiche CF & De Clerck O (2012) Phylogeny and molecular evolution of the green algae. Critical Reviews in Plant Sciences 31: 1-46. pdf
The green lineage (Viridiplantae) comprises the green algae and their descendants the land plants, and is one of the major groups of oxygenic photosynthetic eukaryotes. Current hypotheses posit the early divergence of two discrete clades from an ancestral green flagellate. One clade, the Chlorophyta, comprises the early diverging prasinophytes, which gave rise to the core chlorophytes. The other clade, the Streptophyta, includes the charophyte green algae from which the land plants evolved. Multi-marker and genome scale phylogenetic studies have greatly improved our understanding of broad-scale relationships of the green lineage, yet many questions persist, including the branching orders of the prasinophyte lineages, the relationships among core chlorophyte clades (Chlorodendrophyceae, Ulvophyceae, Trebouxiophyceae and Chlorophyceae), and the relationships among the streptophytes. Current phylogenetic hypotheses provide an evolutionary framework for molecular evolutionary studies and comparative genomics. This review summarizes our current understanding of organelle genome evolution in the green algae, genomic insights into the ecology of oceanic picoplanktonic prasinophytes, molecular mechanisms underlying the evolution of complexity in volvocine green algae, and the evolution of genetic codes and the translational apparatus in green seaweeds. Finally, we discuss molecular evolution in the streptophyte lineage, emphasizing the genetic facilitation of land plant origins.
I. The nature and origins of green algae and land plants
II. Green lineage relationships
A. Morphology, ultrastructure and molecules
B. Phylogeny of the green lineage
1. Two main lineages: Chlorophyta and Streptophyta
2. Early diverging Chlorophyta: the prasinophytes
3. The core Chlorophyta: ecological and morphological diversification
3.1. Radiation of the Ulvophyceae, Trebouxiophyceae and Chlorophyceae
3.2. Trebouxiophyceae
3.3. Chlorophyceae
3.4. Ulvophyceae
4. Streptophyta: charophyte green algae and the origin of land plants
4.1. The early diverging charophytes
4.2. The closest living relative of the land plants
III. Spread of green genes in other eukaryotes
IV. Green algal evolution: insights from genes and genomes
A. Organelle genome evolution
B. Ecology and molecular evolution of oceanic picoplanktonic prasinophytes
C. Genomic insights into the evolution of complexity in volvocine green algae
D. Genetic codes and the translational apparatus in green seaweeds
E. Molecular evolution in the Streptophyta and the origin of land plants
V. Conclusions and perspectives

I. The nature and origins of green algae and land plants
The green lineage or Viridiplantae* includes the green algae and land plants, and is one of the major groups of oxygenic photosynthetic eukaryotes. Green algae are diverse and ubiquitous in aquatic and some terrestrial habitats, and they have played a crucial role in the global ecosystem for hundreds of millions of years (Falkowski et al., 2004; O’Kelly, 2007). The evolution of land plants from a green algal ancestor was a key event in the history of life and has led to dramatic changes in the earth’s environment, initiating the development of the entire terrestrial ecosystem (Kenrick & Crane, 1997).
The green lineage originated following an endosymbiotic event in which a heterotrophic eukaryotic host cell captured a cyanobacterium that became stably integrated and ultimately turned into a plastid (Archibald, 2009; Keeling, 2010). This primary endosymbiosis, which likely happened between 1 and 1.5 billion years ago (Hedges et al., 2004; Yoon et al., 2004), marked the origin of the earliest oxygenic photosynthetic eukaryotes. The subsequent diversification of this primary plastid-containing eukaryote gave rise to the green lineage, as well as the red algae and the glaucophytes. From this starting point, photosynthesis spread widely among diverse eukaryotic protists via secondary and tertiary endosymbioses, which involved captures of either green or red algae by non-photosynthetic protists (Keeling, 2010). Secondary endosymbioses involving green algae as the autotrophic partner have given rise to three groups of algae: the chlorarachniophytes, the photosynthetic euglenids and the “green” dinoflagellates (see section III. Spread of green genes in other eukaryotes). The other eukaryotic algal groups, the cryptophytes, haptophytes, photosynthetic stramenopiles (e.g., diatoms, chrysophytes and brown seaweeds) and dinoflagellates, have acquired plastids from a red algal ancestor, either by a single or multiple endosymbiotic events (Archibald, 2009; Bodyl et al., 2009; Baurain et al., 2010).
The green lineage is ancient, and dating its origin has been a difficult task because of the sparse fossil record of the group. The earliest fossils attributed to green algae date from the Precambrian (ca. 1200 mya) (Tappan, 1980; Knoll, 2003). The nature of these early fossils, however, remains controversial (e.g., Cavalier-Smith, 2006). The resistant outer walls of prasinophyte cysts (phycomata) are well preserved in fossil deposits and especially abundant and diverse in the Paleozoic era (ca. 250-540 mya) (Parke et al., 1978; Tappan, 1980; Colbath, 1983). A filamentous fossil (Proterocladus) from middle Neoproterozoic deposits (ca. 750 mya) has been attributed to siphonocladous green algae (Cladophorales) (Butterfield et al., 1994; Butterfield, 2009), while the oldest reliable records of the siphonous seaweeds (Bryopsidales, Dasycladales) and stoneworts (Charophyceae) are from the Paleozoic (Hall & Delwiche, 2007; Verbruggen et al., 2009a). The earliest land plant fossils are Mid-Ordovician in age (ca. 460 mya) (Kenrick & Crane, 1997; Steemans et al., 2009). Molecular clock analyses have estimated the origin of the green lineage between 700 and 1500 mya (Douzery et al., 2004; Hedges et al., 2004; Berney & Pawlowski, 2006; Roger & Hug, 2006; Herron et al., 2009). These estimates are sensitive to differences in methodology and interpretation of fossils and tend to yield older dates than are well supported by the fossil record. This could be attributable to miscalibration of the molecular clock estimates or to taphonomic bias and the difficulty of interpreting fossils with no modern exemplars. Molecular phylogenetic evidence has provided a substantially improved understanding of the relationships among major lineages. Reconstruction of ancestral character states could assist in the reinterpretation of known specimens of uncertain affinity, and this, combined with continued paleontological investigation, holds out hope for reconciliation of molecular and fossil evidence.
Green algae are characterized by a number of distinct features, many of which are also shared with the land plants (van den Hoek et al., 1995; Graham et al., 2009). The chloroplasts are enclosed by a double membrane with thylakoids grouped in lamellae, and contain chlorophyll a and b along with a set of accessory pigments such as carotenes and xanthophylls. Pyrenoids, when present, are embedded within the chloroplast and are surrounded by starch, which is the main reserve polysaccharide. Most green algae have firm cell walls with a fiber matrix generally composed of cellulose. The flagellate cells are isokont, which means that the two flagella are similar in structure, although they may differ in length. The flagellar transition zone (i.e., the region between the flagellum and its basal body) is typically characterized by a stellate structure, which is a nine-pointed star, visible in cross section using an electron microscope, linking nine pairs of microtubules (Melkonian, 1984).
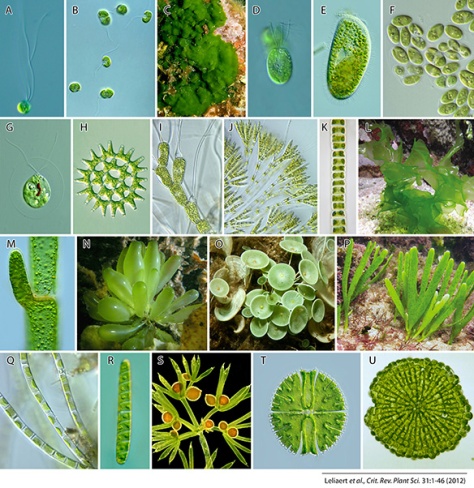
Despite their many unifying features, green algae exhibit a remarkable variation in morphology and ecology reflecting their evolutionary diversification. Morphological diversity ranges from the smallest known free-living eukaryote, Ostreococcus tauri, to large, multicellular life forms (Fig. 1). Green algae are especially abundant and diverse in freshwater environments, including lakes, ponds, streams and wetlands (John et al., 2002; Wehr & Sheath, 2003), where they may form nuisance blooms under conditions of nutrient pollution (Malkin et al., 2010). Only two green algal groups are well represented in marine environments. The green seaweeds (Ulvophyceae) abound in coastal habitats. Some green seaweeds (mainly Ulva) can form extensive, free-floating coastal blooms, called ‘green tides’ (Leliaert et al., 2009c); others, like Caulerpa and Codium are notorious for their invasive nature (Meinesz & Hesse, 1991; Jousson et al., 2000; Lapointe et al., 2005). The prasinophytes are planktonic green algae that occur mainly in oceanic environments and are especially abundant in more eutrophic, near-shore waters, where they can form monospecific blooms (O’Kelly et al., 2003; Not et al., 2004). Embryophytes have dominated the terrestrial environment since the late Ordovician, and some have become secondarily adapted to aquatic environments, including holoaquatic marine species that form extensive beds of seagrass. Several green algae have adapted to highly specialised or extreme environments, such as hot or cold deserts (Lewis & Lewis, 2005; De Wever et al., 2009; Schmidt et al., 2011), hypersaline habitats (Vinogradova & Darienko, 2008), acidic waters with extreme concentrations of heavy metals (Zettler et al., 2002), marine deep waters (Zechman et al., 2010) and deep-sea hydrothermal vents (Edgcomb et al., 2002). Some green algal groups, i.e., Trentepohliales, are exclusively terrestrial and never found in aquatic environments (López-Bautista et al., 2006). Several lineages have engaged in symbiosis with a diverse range of eukaryotes, including fungi to form lichens, ciliates, foraminifers, cnidarians, molluscs (nudibranchs and giant clams) and vertebrates (Friedl & Bhattacharya, 2002; Lewis & Muller-Parker, 2004; Kovacevic et al., 2010; Kerney et al., 2011). Others have evolved an obligate heterotrophic life style as parasites or free-living species (Joubert & Rijkenberg, 1971; Rumpf et al., 1996; Huss et al., 1999; Nedelcu, 2001). The heterotrophic green alga Prototheca, which grows in sewage and soil, can cause infections in humans and animals known as protothecosis (Sudman, 1974).
Several green algae serve as model systems or are of economic importance. Melvin Calvin used cultures of Chlorella to elucidate the light-independent reactions of photosynthesis, now known as the Calvin cycle (Calvin & Benson, 1948). Transplant experiments with the giant-celled Acetabularia, conducted by Joachim Hämmerling, demonstrated that the nucleus of a cell contains the genetic information that directs cellular development, and postulated the existence of messenger RNA before its structure was determined (Hämmerling, 1953). Acetabularia, along with other giant-celled green algae (Valonia, Chara and Nitella), has also served as an experimental organism for electro-physiological research and studies of cell morphogenesis (Menzel, 1994; Mandoli, 1998; Shepherd et al., 2004; Bisson et al., 2006; Mine et al., 2008). The charophyte Mougeotia played a key role in outlining the role of phytochrome in plant development (Winands & Wagner, 1996). The biochemistry and physiology of the unicellular, halophilic Dunaliella salina have been studied in great detail. This alga is among the most industrially important microalgae because it can produce massive amounts of β-carotene that can be collected for commercial purposes, and because of its potential as a feedstock for biofuels production (Oren, 2005; Gouveia & Oliveira, 2009; Tafresh & Shariati, 2009). The unicellular flagellate Chlamydomonas reinhardtii has long been used as a model system for studying photosynthesis, chloroplast biogenesis, flagellar assembly and function, cell-cell recognition, circadian rhythm and cell cycle control because of its well-defined genetics, and the development of efficient methods for nuclear and chloroplast transformation (Rochaix, 1995; Harris, 2001; Grossman et al., 2003; Breton & Kay, 2006). The colonial green alga Volvox has served as a model for the evolution of multicellularity, cell differentiation, and colony motility (Kirk, 1998; Kirk, 2003; Herron & Michod, 2008; Herron et al., 2009).
Analysis of the complete nuclear genome sequence of C. reinhardtii greatly advanced our understanding of ancient eukaryotic features such as the function and biogenesis of chloroplasts, flagella and eyespots, and regulation of photosynthesis (Merchant et al., 2007; Kreimer, 2009; Peers et al., 2009). Genome data are rapidly accumulating and to date seven complete green algal genomes have been sequenced: the prasinophytes Ostreococcus tauri (Derelle et al., 2006), O. lucimarinus (Palenik et al., 2007) and two isolates of Micromonas pusilla (Worden et al., 2009), the chlorophytes C. reinhardtii (Merchant et al., 2007) and Volvox carteri (Prochnik et al., 2010), and the trebouxiophyte Chlorella variabilis (Blanc et al., 2010). Several other genome projects are ongoing, including the complete sequencing of Coccomyxa, Dunaliella, Bathycoccus, Botryococcus and additional Ostreococcus and Micromonas strains (Tirichine & Bowler, 2011). These data provide a great resource for in-depth analysis of genome organization and the processes of eukaryotic genome evolution. In addition, green algal genomes are important sources of information for the evolutionary origins of plant traits because of their evolutionary relationship to land plants.
Reconstruction of the phylogenetic relationships among green plants is essential to identifying the innovations underlying the diversity of green algae and land plants. Molecular phylogenetics has dramatically reshaped our views of green algal relationships and evolution. This review summarizes the current understanding of green algal phylogeny, focusing primarily on relationships among the major lineages of green algae, which are usually classified as divisions, classes and orders. Current phylogenetic hypotheses have provided an evolutionary framework for molecular evolutionary studies and comparative genomics. In this review, we highlight a number of topics, including the evolution of organellar genomes, ecology and molecular evolution of marine picoplanktonic prasinophytes, genomic insights into the evolution of complexity in volvocine green algae, molecular evolution in the green seaweeds, and molecular evolution in the streptophyte green algae and the origin of land plants.
* Various names have been proposed for the lineage comprising the green algae and land plants: “Viridiplantae” or “Viridaeplantae” (Cavalier-Smith, 1981, 1998), “Chlorobiota” or “Chlorobionta” (Jeffrey, 1971, 1982), “Chloroplastida” (Adl et al. 2005), or simply “green plants” (Sluiman et al. 1983) or “green lineage”.
II. Green lineage relationships
A. Morphology, ultrastructure and molecules
Early hypotheses of green algal phylogeny were based on the concept that evolution follows trends in levels of morphological complexity (Fritsch, 1935; Fott, 1971). Unicellular flagellates were believed to have initially evolved into non-motile unicells (coccoid) and loose packets of cells (sarcinoid), followed by various multicellular forms and siphonous algae. This hierarchy reflected the view that the morphologies that are organized in two- and three-dimensional space require more elaborate developmental controls, and hence would be expected to appear later in an evolutionary sequence. In this view the land plants were derived from more complex, filamentous green algae (Blackman, 1900; Pascher, 1914).
A large amount of new information was gathered in the 1970s and 80s, mainly from investigations of the fine structures of green algal cells and life cycles (Round, 1984). These data led to a thorough re-evaluation of evolutionary relationships and a revised classification of green algae, primarily based on flagellar ultrastructure and processes of mitosis and cell division (Picket-Heaps & Marchant, 1972; Melkonian, 1982; Mattox & Stewart, 1984; Melkonian, 1984; O’Kelly & Floyd, 1984a; O’Kelly & Floyd, 1984b; van den Hoek et al., 1988). These features, which apply to most (but not all) green algae, were believed to accurately reflect phylogenetic relationships because of their involvement in fundamental processes of cell replication and cell motility, and thus less liable to convergent evolution than gross morphological traits. Ultrastructure-based phylogenetic hypotheses posited an early diversification of flagellate unicells, resulting in a multitude of ancient lineages of flagellates, some of which then evolved into more complex green algae. Although ultrastructural data have laid the foundations for a natural classification of the green algae, analyses of these data have not resolved the phylogenetic relationships among the main green algal lineages.
The introduction of molecular phylogenetic methods provided a new framework for reconstructing the evolutionary history of the green lineage. Analyses of DNA sequence data took a start in the mid 1980’s with the first phylogenies of green plants inferred from 5.8S nrDNA sequences (Hori et al., 1985; Hori & Osawa, 1987), soon followed by 18S and 28S nrDNA sequence analyses (Gunderson et al., 1987; Perasso et al., 1989; Buchheim et al., 1990; Zechman et al., 1990; Chapman et al., 1991; Mishler et al., 1992; Chapman et al., 1998). Ribosomal sequences were chosen at the time because enough RNA could be obtained for direct sequencing and (later) because regions of the gene were conserved enough to make universal primers. Ribosomal RNA is also found in all known living cells, and although it is typically present in multiple copies in the genome, concerted evolution homogenizes sequence diversity, and consequently is assumed to reduce the risk of sequencing paralogs. Early molecular studies were pre-PCR and hence involved laboratory cloning and generally few taxa. Since 1990 rapid advancements in techniques in molecular biology (e.g., the utilization of PCR) and bioinformatics made it possible to generate and analyze larger datasets. Nuclear-encoded 18S rDNA sequences have been, until recently, the primary source of data for inferring phylogenetic relationships among green algae (Pröschold & Leliaert, 2007), supplemented by 28S rDNA (e.g., Buchheim et al., 2001; Shoup & Lewis, 2003), actin (An et al., 1999) and the chloroplast genes rbcL, tufA and atpB (e.g., Daugbjerg et al., 1994; Daugbjerg et al., 1995; McCourt et al., 2000; Hayden & Waaland, 2002; Nozaki et al., 2003; Zechman, 2003; Rindi et al., 2007).
These initial molecular phylogenetic investigations have generally corroborated the ultrastructure-based higher-level classification of the green algae, but have also revised the circumscriptions of several lineages (McCourt, 1995). However, analyses of individual genes have only partly resolved the relationships among the main green algal lineages. It is now clear that a large number of genes from many species must be analysed to arrive at a reliable phylogenetic resolution for an ancient group such as the green algae (Philippe & Telford, 2006). These datasets have mainly involved concatenated sequences of protein-coding genes that are shared among green algal chloroplast genomes. To date, 26 complete green algal plastid genomes have been sequenced and assembled (Wakasugi et al., 1997; Turmel et al., 1999b; Lemieux et al., 2000; Maul et al., 2002; Turmel et al., 2002b; Pombert et al., 2005; Turmel et al., 2005; Bélanger et al., 2006; de Cambiaire et al., 2006; Pombert et al., 2006; Turmel et al., 2006a; de Cambiaire et al., 2007; Lemieux et al., 2007; Robbens et al., 2007a; Brouard et al., 2008; Turmel et al., 2008; Turmel et al., 2009a; Turmel et al., 2009b; Zuccarello et al., 2009; Brouard et al., 2010; Brouard et al., 2011), in addition to more than 30 angiosperm plastid genomes (Soltis et al., 2009). Chloroplast genomes are particularly useful for phylogenetic reconstruction because of their relatively high and condensed gene content, in comparison to nuclear genomes. Furthermore, in contrast to many nuclear genes that are multi-copy in nature, which can confound phylogenetic reconstruction, organellar genes are typically single-copy and do not pose these problems. Only recently have multi-gene analyses of nuclear genes been carried out (e.g., Rodríguez-Ezpeleta et al., 2007; Cocquyt et al., 2010b; Finet et al., 2010).
A persistent problem in many of the multi-gene phylogenetic investigations thus far is sparse and incomplete taxon sampling, which may result in systematic errors in phylogenetic reconstruction. For this reason, phylogenetic hypotheses derived from such studies are preferably supported by independent data, such as structural genomic features like gene content, gene order, gene structure or intron distribution (Lemieux et al., 2007). But ideally, these systematic errors would be overcome by increasing taxon sampling. Recent advances in high-throughput DNA sequencing, including Roche-454 and Illumina-Solexa (Shendure & Ji, 2008; Metzker, 2010) facilitate rapid sequencing of organellar genomes, transcriptomes and entire nuclear genomes. This fast accumulation of genomic data in conjunction with new developments in phylogenetic inference techniques is creating unprecedented research opportunities. The reconstruction of large-scale multi-gene phylogenies and studies of the molecular mechanisms underlying this diversity is now a short-term feasible prospect.
B. Phylogeny of the green lineage
1. Two main lineages: Chlorophyta and Streptophyta
Current hypotheses on green algal evolution posit the early divergence of two discrete lineages: the Chlorophyta and Streptophyta (Fig. 2) (Picket-Heaps & Marchant, 1972; Bremer, 1985; Lemieux et al., 2007). The Chlorophyta includes the majority of described species of green algae. The Streptophyta are comprised of the charophytes, a paraphyletic assemblage of freshwater algae, and the land plants.
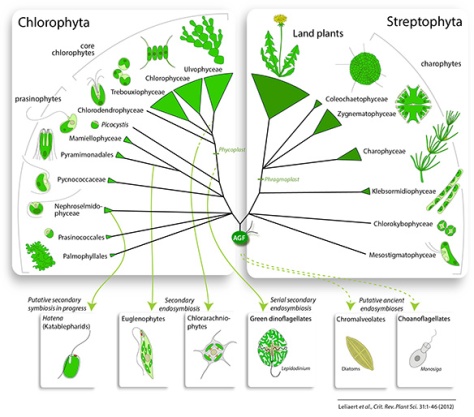
–
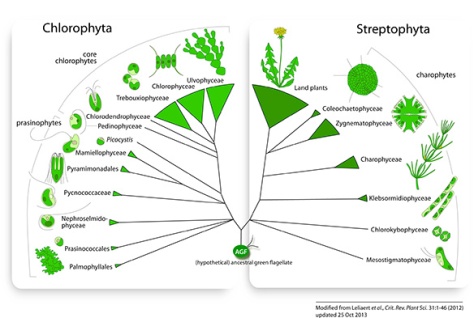
– 2 May 2012: Marin (2012) provided evidence for an independent class Pedinophyceae, sister to all phycoplast-containing core chlorophytes (Chlorodendrophyceae, Trebouxiophyceae, Ulvophyceae and Chlorophyceae).
– 8 Aug 2012: Re-analysis of the dataset of Finet et al. (2010) showed that land plants are sister to a group composed of Zygnematophyceae and Coleochaetophyceae (Laurin-Lemay et al. 2012, Finet et al. 2012)
Charophytes range in morphology from unicellular to complex multicellular organisms (Fig. 1Q-U) and primarily occur in freshwater and, to a lesser extent, terrestrial habitats. They display a number of ultrastructural and biochemical traits that are shared with land plants but not with the Chlorophyta: motile cells (when present) with two subapically inserted flagella and an asymmetrical flagellar apparatus that contains a distinctive multilayered structure (MLS) and parallel basal bodies, open mitosis and persistent mitotic spindle, a phragmoplast-type cytokinesis (in many species), and the presence of several unique enzyme systems like Cu/Zn superoxide dismutase, class I aldolases and glycolate oxydase (Frederick et al., 1973; Mattox & Stewart, 1984; de Jesus et al., 1989; Jacobshagen & Schnarrenberger, 1990; Graham & Kaneko, 1991). The Streptophyta are more broadly characterized by discrete molecular features, such as a GapA/B gene duplication in the common ancestor of the clade (Petersen et al., 2006), while other molecular characteristics, such as polyadenylation signals, show a more complex phylogenetic distribution (Wodniok et al., 2007).
Members of the Chlorophyta are common inhabitants of marine, freshwater and terrestrial environments and exhibit a remarkable morphological diversity (Fig. 1A-P). The flagellar apparatus is typically characterized by a symmetrical cruciate root system wherein rootlets of variable (X) numbers of microtubules alternate with rootlets composed of two microtubules to form a “X-2-X-2” arrangement (Moestrup, 1978). The orientation of the flagellar root system has served as an important character for defining the main groups of Chlorophyta (Mattox & Stewart, 1984; O’Kelly & Floyd, 1984b). When viewed from the anterior (flagellar) end of the cell, the flagellar basal bodies and rootlets can have a perfect cruciate pattern (i.e., with basal bodies directly opposed, DO) or they are offset in a counter-clockwise (CCW) or clockwise (CW) position (for a detailed overview, see van den Hoek et al., 1995; Graham et al., 2009). Traditionally, four classes are recognized within the Chlorophyta: the freshwater or terrestrial Trebouxiophyceae and Chlorophyceae, the coastal Ulvophyceae, and the unicellular, predominantly marine planktonic Prasinophyceae.
Resolution of the earliest green algal divergences often remains elusive because of the antiquity of the green lineage and the rapidity of the early evolutionary radiations (McCourt, 1995). Figures 2 and 3 illustrate our current understanding of green algal relationships. The topology presented is a somewhat conservative interpretation of available data with polytomies representing uncertainties or conflicts between different studies. The evidence supporting these relationships is discussed below on a clade-by-clade basis.
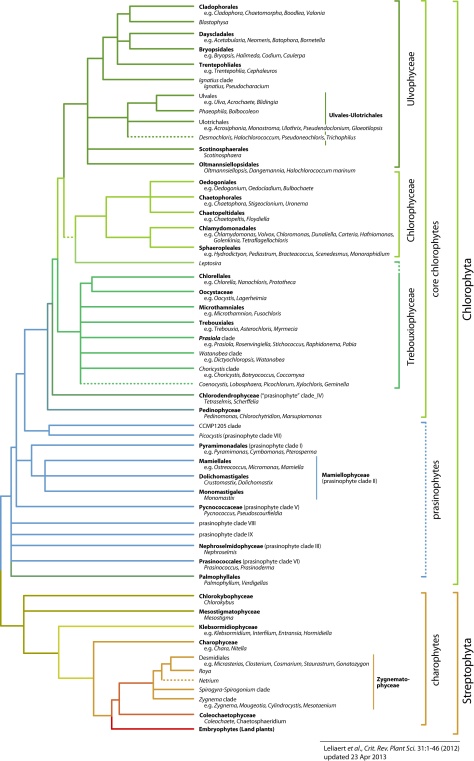
2. Early diverging Chlorophyta: the prasinophytes
The Prasinophyceae originally comprised motile unicells with organic body scales (Mattox & Stewart, 1984). The ultrastructure of the body scales and the flagellar apparatus, along with cell division and mitotic features, have long served as the main characters to distinguish between the major groups of Prasinophyceae (Melkonian, 1990b; Sym & Pienaar, 1993; Becker et al., 1994; Marin & Melkonian, 1994). Molecular data have provided clear evidence that the class forms a paraphyletic assemblage of early diverging lineages (Kantz et al., 1990; Steinkötter et al., 1994; Nakayama et al., 1998), which is reflected by the wide diversity of cell shapes, body scale morphologies, flagellar behaviour, mitotic and cytokinetic processes, and biochemical features (Sym & Pienaar, 1993; Latasa et al., 2004).
Based on the early diverging nature of the Prasinophyceae, the ancestor of the green lineage is believed to be a flagellate unicell. However, the nature of this ancestral green flagellate (AGF) remains tentative. Simple flagellates have been proposed to most closely resemble the AGF (Moestrup, 1991). Alternatively, the food uptake apparatus of some complex mixotrophic prasinophytes (e.g. Cymbomonas) has been interpreted as a feature inherited from a phagotrophic ancestor of the green lineage that was subsequently lost in most green algae (Moestrup et al., 2003; O’Kelly, 2007; Turmel et al., 2009a).
The circumscription of the prasinophytes has changed considerably with the discovery of several new taxa and environmental sequencing (Hasegawa et al., 1996; Nakayama et al., 1998; Fawley et al., 2000; Zingone et al., 2002; Guillou et al., 2004; Vaulot et al., 2008; Viprey et al., 2008; Lepère et al., 2009; Shi et al., 2009). The prasinophytes, as presently conceived, include a heterogeneous assemblage of scaled (e.g., Nephroselmis, Pyramimonas and Mamiella) and naked (e.g., Ostreococcus, Micromonas and Scourfieldia) unicells with diverse morphologies with one to eight flagella. In addition, several coccoid forms have been found, which are distributed among at least four clades (Fawley et al., 2000; Guillou et al., 2004; Jouenne et al., 2011). Prasinophytes are predominantly marine planktonic but also include several freshwater representatives (Marin & Melkonian, 2010). Although sexual reproduction has rarely been documented in prasinophytes (but see Suda et al., 2004), some members (e.g., Pyramimonadales) produce resistant cysts (phycomata) containing two chloroplasts, indicative of sexual reproduction (Moestrup et al., 2003). Sexual reproduction has also been implied in Micromonas and Ostreococcus based on the presence of meiosis-specific and sex-related genes in the four sequenced genomes (see below) (Derelle et al., 2006; Worden et al., 2009; Grimsley et al., 2010).
The paraphyletic nature of prasinophytes has important consequences for understanding early green algal evolution, and a well-resolved phylogeny has the potential to shed light on the nature of the common ancestor of the Chlorophyta and on the origin of the core chlorophytes (Turmel et al., 2009a). Multi-gene phylogenetic analyses are just beginning to shed light on these early divergences. To date five chloroplast genomes from prasinophyte representatives of four clades (Mamiellophyceae, Nephroselmidophyceae, Pycnococcaceae and Pyramimonadales) have been sequenced, in addition to Mesostigma, a former member of the prasinophytes that is now known to be an early-branching streptophyte (Turmel et al., 1999b; Lemieux et al., 2000; Robbens et al., 2007a; Turmel et al., 2009a).
The Mamiellophyceae is the largest lineage of prasinophytes and unites the morphologically and ecologically diverse Mamiellales, the Monomastigales (Monomastix) and Dolichomastigales (Dolichomastix and Crustomastix) (Marin & Melkonian, 2010). The Mamiellales includes marine and freshwater flagellates with one or two laterally inserted flagella, as well as coccoid forms, with or without body scales. They include some of the smallest eukaryotes known (e.g., Ostreococcus, Micromonas), which are prominent in the oceanic picoplankton (Courties et al., 1994; O’Kelly et al., 2003; Guillou et al., 2004; Not et al., 2004). The clade is further characterized by prasinoxanthin, a pigment that is also found in some other clades, such as the Pycnococcaceae (Zingone et al., 2002; Latasa et al., 2004). The phylogenetic affinity of the freshwater, uniflagellate Monomastix has long been uncertain because it lacks scales and has atypical surface structures (Manton, 1967; Melkonian, 1990b; Sym & Pienaar, 1993). Analyses of 18S sequence and chloroplast genome data clearly show that Monomastix is sister to the Mamiellales (Turmel et al., 2009a). The biflagellate Dolichomastigales are also morphologically distinct: Dolichomastix has atypical scales and Crustomastix has a specialized cell covering rather than the more typical body scales (Nakayama et al., 2000; Zingone et al., 2002). Their phylogenetic placement in Mamiellophyceae has been demonstrated by 18S data (Zingone et al., 2002). The relationships among the Mamiellales, Monomastigales and Dolichomastigales, however, have not been resolved.
Chloroplast phylogenomic analyses resolved a sister relationship between the Mamiellophyceae and the Pyramimonadales (Turmel et al., 2009a), a clade of relatively large quadriflagellates with diverse and complex body scale coverings that occur in marine and freshwater habitats (Melkonian, 1990b). As mentioned above, some Pyramimonadales (e.g. Cymbomonas) are unique among green algae in possessing a food uptake apparatus (Moestrup et al., 2003).
The Pycnococcaceae is a small clade containing the marine biflagellate Pseudoscourfieldia and the naked coccoid Pycnococcus. Some 18S studies have related this clade with the Nephroselmidophyceae (although never with strong support; Fawley et al., 2000; Guillou et al., 2004), which is corroborated by some morphological similarities (e.g., Nephroselmis and Pseudoscourfieldia both have two unequal flagella and two body and flagellar scale layers) (Nakayama et al., 2007). However, chloroplast multi-gene analyses do not support a relationship between the two clades and have identified the Nephroselmidophyceae as an early diverging prasinophyte lineage, while the phylogenetic position of the Pycnococcaceae remains equivocal (Turmel et al., 2009a).
Environmental sequencing of photosynthetic picoeukaryote communities in the Mediterranean Sea and SE Pacific Ocean has identified two additional prasinophyte clades (clades VIII and IX) (Viprey et al., 2008; Lepère et al., 2009; Shi et al., 2009). However, the nature and phylogenetic affinities of these clades remain elusive.
Zechman et al. (2010) provided evidence for another early diverging lineage of green algae, the Palmophyllales. This lineage includes Palmophyllum and Verdigellas, green seaweeds that thrive in deepwater and other dimly lit, benthic marine habitats (Womersley, 1984; Nelson & Ryan, 1986; Ballantine & Norris, 1994). Palmophyllales feature a unique type of multicellularity, forming firm, well-defined macroscopic thalli composed of isolated spherical cells in a gelatinous matrix (Pueschel et al., 1997). Analysis of two plastid genes (atpB and rbcL) placed the Palmophyllales as the sister clade to all other Chlorophyta while 18S phylogenetic analysis allied the Palmophyllales with the Prasinococcales clade. This clade includes the coccoid prasinophytes Prasinococcus and Prasinoderma, and it has also been shown to form an early diverging prasinophyte clade based on 18S data (Guillou et al., 2004; Turmel et al., 2009a). The possible relationship between the Palmophyllales and Prasinococcales is supported by a number of shared cytological features and similarities in cell division (O’Kelly, 1988; Hasegawa et al., 1996; Zechman et al., 2010; Jouenne et al., 2011).
Sequencing of environmental samples and cultures has identified a clade of coccoid prasinophytes (CCMP1205 clade) that, together with the saline lake dwelling coccoid Picocystis, emerges as a sister lineage to the core chlorophytes, although strong support for this relationship is still lacking (Guillou et al., 2004; Marin & Melkonian, 2010).
3. The core Chlorophyta: ecological and morphological diversification
The prasinophytes have given rise to the morphologically and ecologically diverse core chlorophytes. This group includes the early diverging Chlorodendrophyceae, and three major classes: Ulvophyceae, Trebouxiophyceae and Chlorophyceae (UTC). The core chlorophytes are characterized by a new mode of cell division that is mediated by a phycoplast (i.e., a system of microtubules that develops parallel to the plane of nuclear division), which was subsequently lost in the Ulvophyceae.
The Chlorodendrophyceae is a small clade uniting the marine or freshwater scaly quadriflagellates Tetraselmis and Scherffelia (Guillou et al., 2004). These unicells were traditionally regarded as members of the Prasinophyceae but they share several features with the UTC clades, including closed mitosis and a phycoplast (Mattox & Stewart, 1984). The close relationship with the UTC classes has been confirmed by 18S and multi-gene phylogenetic data (Massjuk, 2006; Cocquyt et al., 2010b).
3.1. Radiation of the Ulvophyceae, Trebouxiophyceae and Chlorophyceae
The UTC classes are species-rich and morphologically and ecologically diverse. Ecophysiological adaptations have likely led to the success of the Chlorophyceae and Trebouxiophyceae in freshwater and terrestrial environments, while the Ulvophyceae mainly diversified in coastal ecosystems. Marine versus freshwater lifestyles also coincide with differentiations in life histories. Whereas the marine Ulvophyceae mainly have life cycles involving an alternation between a free-living haploid, gametophytic and a free-living diploid, sporophytic multicellular generation, most freshwater green algae have a haploid vegetative phase and a single-celled, often dormant zygote as the diploid stage. In terrestrial members of the core chlorophytes, sexual reproduction has rarely been documented (Rindi, 2011).
Resolving the phylogenetic relationships among and within the UTC classes can provide important insights into the evolution of these ecophysiological, life history and morphological traits. Molecular phylogenetic studies based on 18S sequences (Zechman et al., 1990; López-Bautista & Chapman, 2003; Watanabe & Nakayama, 2007), chloroplast and mitochondrial multigene data (Pombert et al., 2004; Pombert et al., 2005; Turmel et al., 2009b) have yielded ambivalent results. All possible relationships have been hypothesized, depending on interpretation of ultrastructural characters, gene and taxon sampling, and phylogenetic methods used. Furthermore, the monophyly of the Trebouxiophyceae and Ulvophyceae has not been unequivocally demonstrated (Pröschold & Leliaert, 2007; Zuccarello et al., 2009). The unstable relationships exhibited among these three classes likely result from their antiquity and the short time span over which they diverged from one another (O’Kelly, 2007; Cocquyt et al., 2010b). The fossil record indicates the presence of the classes in the mid-Neoproterozoic and molecular clock estimates situate the UTC divergence in the early Neoproterozoic (Butterfield et al., 1994; Douzery et al., 2004; Herron et al., 2009).
Some early 18S phylogenies showed a sister relationship between Chlorophyceae and Trebouxiophyceae (e.g., Krienitz et al., 2001), while more recent studies with increased taxon sampling revealed a sister relationship between the Chlorophyceae and Ulvophyceae (e.g., Friedl & O’Kelly, 2002; Watanabe & Nakayama, 2007; De Wever et al., 2009). Chloroplast multi-gene phylogenetic analyses have generally supported a sister relationship between Ulvophyceae and Trebouxiophyceae (Pombert et al., 2005; Turmel et al., 2009b), although some analyses based on nucleotide sequences suggest a sister relationship between Ulvophyceae and Chlorophyceae (Turmel et al., 2008; Turmel et al., 2009a). The latter topology is also supported by a mitochondrial multi-gene analysis (Pombert et al., 2004), a phylogenetic analysis of eight nuclear and two plastid genes (Cocquyt et al., 2010b), and structural chloroplast genome data (Pombert et al., 2005; Turmel et al., 2009b). Ultrastructural data have been interpreted as either providing supports for a sister relationship between Chlorophyceae and Trebouxiophyceae or between Trebouxiophyceae and Ulvophyceae. A relationship between Chlorophyceae and Trebouxiophyceae has been suggested based on the shared presence of a non-persistent mitotic spindle (Mattox & Stewart, 1984). A relationship between Trebouxiophyceae and Ulvophyceae was proposed based on a counter-clockwise orientation of the flagellar apparatus (Sluiman, 1989). In contrast, molecular data generally support an early diverging Trebouxiophyceae. This would imply the ancestral status of a CCW orientation of the flagellar basal bodies, which evolved to a DO and CW orientation in the Chlorophyceae. This interpretation is congruent with a CCW flagellar root system in the Chlorodendrophyceae.
The genus-level systematics of the core chlorophytes has been profoundly misled by convergent evolution toward reduced morphologies like unicells or simple filaments, and many genera defined on these features have been shown to be polyphyletic (Lewis & McCourt, 2004). For example, strains conforming to the morphological circumscription of the genera Neochloris, Characium and Planophila have been found in all three classes (Watanabe & Floyd, 1989; Lewis et al., 1992; Watanabe et al., 2000; Friedl & O’Kelly, 2002). Similarly, species of the coccoid genus Chlorococcum and the filamentous genus Uronema were present in the Chlorophyceae as well as the Ulvophyceae (Watanabe et al., 2001; Krienitz et al., 2003; Leliaert et al., 2009a). Species of Chlorella were shown to be distributed between the Chlorophyceae and Trebouxiophyceae (Huss & Sogin, 1990; Huss et al., 1999; Luo et al., 2010).
3.2. Trebouxiophyceae
The Trebouxiophyceae was originally defined (as Pleurastrophyceae) based on ultrastructural features (CCW flagellar apparatus orientation, non-persistent metacentric spindle, and phycoplast-mediated cytokinesis) (Mattox & Stewart, 1984) and its circumscription was later refined by 18S sequence data (Kantz et al., 1990; Friedl, 1995; Wolf et al., 2003) (see Lewis & McCourt, 2004 for a taxonomic and nomenclatural history of the class). As presently conceived, the class encompasses motile and non-motile unicells, colonies and multicellular filaments or blades from freshwater or terrestrial habitats, with some species penetrating in brackish or marine waters. Several species engage in symbiotic relationships with fungi to form lichens (Friedl & Büdel, 1996; Friedl & Bhattacharya, 2002). Others are photosynthetic endosymbionts in various freshwater and marine protists, invertebrates and plants, including ciliates, mussels, hydra, sea anemones and Ginkgo (“zoochlorellae”) (Karakashian & Karakashian, 1965; Tremouillaux-Guiller et al., 2002; Lewis & Muller-Parker, 2004; Hoshina & Imamura, 2008; Summerer et al., 2008; Letsch et al., 2009). Molecular phylogenetic data have shown that many of these endosymbiotic relationships evolved multiple times independently (Hoshina & Imamura, 2008; Pröschold et al., 2011). A number of trebouxiophytes have lost photosynthetic capacity and have evolved heterotrophic free-living or highly adapted parasitic lifestyles (e.g., Prototheca and Helicosporidium) (de Koning & Keeling, 2006; Pombert & Keeling, 2010).
Several distinct lineages within the Trebouxiophyceae have been recovered by molecular data, including the Chlorellales, Trebouxiales, Microthamniales, Prasiola-clade and several clades that have not yet received a formal name (Katana et al., 2001; Friedl & O’Kelly, 2002; Krienitz et al., 2003; Zhang et al., 2008; Neustupa et al., 2011). Resolving the branching patterns among these lineages can provide important insights into morphological and ecological evolution, and provide clues about the origins and adaptations of symbiotic and parasitic lifestyles, ultimately leading to obligate heterotrophic lifestyles. Knowledge of trebouxiophyte interrelationships, however, is limited. Single gene (mostly 18S) analyses were not able to resolve the relationships among the main lineages, and some analyses even cast doubt on the monophyly of the group (Krienitz et al., 2003).
Molecular data have revealed extreme polyphyly of morphologically simple genera indicating convergent evolution toward reduced morphology. In addition, polyphyly has been demonstrated in genera with more complex colonial forms (e.g., Dictyosphaerium, Micractinium and Diacanthos) (Krienitz et al., 2010; Pröschold et al., 2010). On the other hand, molecular phylogenies have shown close relationships between highly dissimilar morphologies, demonstrating that evolution of vegetative morphology can be rapid (Lewis & McCourt, 2004).
The Chlorellales mainly includes freshwater and terrestrial coccoid forms and a few marine members (Aslam et al., 2007). The non-photosynthetic, parasitic Prototheca and Helicosporidium are also included in the Chlorellales based on nuclear and chloroplast gene data (Huss & Sogin, 1990; Tartar et al., 2003; Ueno et al., 2003; de Koning et al., 2005; Ueno et al., 2005). Although plastids are not apparent in Helicosporidium cells, molecular evidence indicates that it maintains a functional plastid genome (Tartar et al., 2003; de Koning & Keeling, 2004; Tartar & Boucias, 2004; de Koning & Keeling, 2006). Diversity and phylogenetic relationships within the Chlorellales have been well studied (Huss et al., 2002; Krienitz et al., 2003; Henley et al., 2004; Zhang et al., 2008; Luo et al., 2010; Pazoutova et al., 2010; Pröschold et al., 2010). 18S and chloroplast genome data provided evidence that the Oocystaceae, including the semi-colonial Oocystis and the spiny Lagerheimia, form an early diverging clade within the Chlorellales (Hepperle et al., 2000; Krienitz et al., 2003; Turmel et al., 2009b).
The enigmatic Pedinomonas is likely related to the Chlorellales. This tiny naked uniflagellate has traditionally been placed in a separate order Pedinomonadales or class Pedinophyceae based on some unusual ultrastructural features (Melkonian, 1990a; Moestrup, 1991). However, chloroplast multi-gene analyses and gene linkage data place Pedinomonas firmly within the Chlorellales (Turmel et al., 2009b). This placement is supported by the CCW orientation of the flagellar apparatus but the persistent telophase spindle in Pedinomonas is atypical for the Trebouxiophyceae. It should also be noted that phylogenetic analyses of seven mitochondrial genes have placed Pedinomonas sister to the Chlorophyceae on a long branch, but these analyses may suffer from low taxon sampling and systematic errors in phylogeny reconstruction (Turmel et al., 1999a; Pombert et al., 2004; Pombert & Keeling, 2010). [Update: Analysis of complete nuclear and plastid-encoded rRNA operon sequences resolved the Pedinophyceae as a sister clade to all phycoplast-containing core chlorophytes (Chlorodendrophyceae, Trebouxiophyceae, Ulvophyceae and Chlorophyceae), supporting the recognition of a separate class (Marin 2012) (see updated Figs 2 and 3)]
The Prasiola-clade is the morphologically and ecologically most diverse trebouxiophyte lineage, including unicellular (e.g., Pseudochlorella, Pseudomarvania, Stichococcus), filamentous (e.g., Rosenvingiella, Raphidonema) and blade-like (e.g., Prasiola) forms growing in freshwater, marine and terrestrial habitats (Karsten et al., 2005; Rindi et al., 2005; Rindi et al., 2007; Elias & Neustupa, 2009).
The Trebouxiales is best known as the “lichen algae group” and includes the common phycobiont genera Trebouxia and Asterochloris (Friedl & Bhattacharya, 2002; Blaha et al., 2006; Skaloud & Peksa, 2010). The order also includes several free-living soil algae (e.g., Myrmecia) (Friedl, 1995). The Microthamniales, including the branched filamentous Microthamnion and the coccoid Fusochloris, is thought to be allied with the Trebouxiales (Lewis & McCourt, 2004), but strong support for this relationship is lacking (Sluiman et al., 2008; Neustupa et al., 2009).
Several other distinct clades of trebouxiophytes, composed of taxa with diverse morphologies and ecologies, have been characterized but their phylogenetic affinities to the better-studied lineages remain uncertain. These clades include the Choricystis lineage, comprising Choricystis, Botryococcus and Coccomyxa (Karsten et al., 2005), the Watanabea clade uniting Dictyochloropsis and Watanabea (Karsten et al., 2005), and various other species that form distinct lineages, Lobosphaera, Parietochloris, Coenocystis, Picochlorum and Xylochloris (Lewis et al., 1992; Hanagata & Chihara, 1999; Henley et al., 2004; Karsten et al., 2005; Neustupa et al., 2011).
The phylogenetic placement of the terrestrial, filamentous genus Leptosira is uncertain (Lokhorst & Rongen, 1994). 18S analyses placed this genus within the Trebouxiophyceae on a long branch with uncertain position (Friedl, 1996; De Wever et al., 2009; Neustupa et al., 2011), while chloroplast multi-gene analyses placed Leptosira on a long branch, sister to the Chlorophyceae with moderate support (Turmel et al., 2009a; Turmel et al., 2009b; Zuccarello et al., 2009). Given the fact that Leptosira was the only trebouxiophycean representative outside the Chlorellales in these analyses, this relationship may be the result of systematic error in phylogenetic reconstruction (Turmel et al., 2009b).
3.3. Chlorophyceae
The Chlorophyceae are a large and morphologically diverse group, including non-motile and motile unicells, colonies, branched and unbranched filaments, and blade-like thalli. Reproduction is equally diverse, including various asexual and sexual modes (van den Hoek et al., 1995). Chlorophycean algae are especially abundant in freshwater but also occur in terrestrial habitats. The class is characterized by closed mitosis during cell division, phycoplast-mediated cytokinesis, and diverse configurations of the flagellar apparatus of motile cells.
Molecular phylogenetic analyses have drastically reshaped the classification of the class. Traditional orders and genera were found to be polyphyletic while others were moved to other classes (e.g., Chlorellales) (see Lewis & McCourt, 2004 for a taxonomic history). Molecular and ultrastructural data have identified five major clades (Booton et al., 1998a; Buchheim et al., 2001; Wolf et al., 2002; Wolf et al., 2003; Turmel et al., 2008). The Chlamydomonadales is characterized by a CW flagellar apparatus orientation in biflagellate members, but quadriflagellate representatives may display various other orientations (Nakayama et al., 1996a; Nozaki et al., 2003; Watanabe et al., 2006b). The Sphaeropleales include nonmotile unicells or colonies that produce biflagellate zoospores with a DO basal body configuration (Deason et al., 1991). The Chaetophorales and Chaetopeltidales are both characterized by quadriflagellate motile cells. The orientation of the flagellar apparatus of the Chaetophorales is variable (DO or CW) (Manton, 1964; Melkonian, 1975; Floyd et al., 1980; Watanabe & Floyd, 1989), whereas the Chaetopeltidales have a DO orientation (O’Kelly & Floyd, 1984a; O’Kelly et al., 1994). The Oedogoniales are characterized by zoospores with an anterior ring of numerous flagella (stephanokont) (Pickett-Heaps, 1975). The atypical flagellar apparatus in the Chaetopeltidales, Oedogoniales and Sphaeropleales has hampered homology assessment with flagellar characters occurring in the other two clades (Turmel et al., 2008).
The phylogenetic relationships among the five main clades of Chlorophyceae have proven difficult to resolve. Phylogenetic analyses of nuclear rDNA data have suggested a sister relationship between the Chlamydomonadales and Sphaeropleales, with the Chaetophorales, Oedogoniales, and Chaetopeltidales forming early diverging clades with uncertain interrelationships (Booton et al., 1998a; Buchheim et al., 2001; Shoup & Lewis, 2003; Müller et al., 2004; Alberghina et al., 2006). Chloroplast multi-gene analyses identified two main lineages within the Chlorophyceae: a clade uniting the Chlamydomonadales and Sphaeropleales (CS clade), and a clade uniting the Oedogoniales, Chaetophorales, and Chaetopeltidales (OCC clade) (Turmel et al., 2008; Brouard et al., 2010). This dichotomy was independently supported by molecular signatures in chloroplast genes, such as presence of indels and transspliced introns. Within the OCC clade, the sister relationship observed for the Chaetophorales and Chaetopeltidales has also been supported by rDNA data (Buchheim et al., 2001; Müller et al., 2004; Caisová et al., 2011) and similarities in zoospore structure (O’Kelly et al., 1994) and the occurrence of two trans-spliced group II introns at identical positions (Turmel et al., 2008). These results have strengthened the notion that the ancestor of the Chlorophyceae was likely a quadriflagellate with a DO+DO configuration of the flagellar apparatus, and that the CW configuration evolved independently in the Chlamydomonadales and the Chaetophorales (Buchheim et al., 2001; Turmel et al., 2008) (but see Nozaki et al., 2003, discussed below), and further suggest that the stephanokont zoospores of the Oedogoniales arose from the DO condition (Turmel et al., 2008).
The Chlamydomonadales forms the largest group of Chlorophyceae and has a complex taxonomic history. Molecular phylogenetic analyses of nuclear ribosomal DNA and chloroplast sequences have profoundly changed the concept of the class, which now includes a diverse range of taxa formally placed in the Dunaliellales, Chlorococcales, Tetrasporales, Chlorosarcinales, Volvocales and Chaetophorales (Lewis et al., 1992; Wilcox et al., 1992; Buchheim et al., 1994; Nakayama et al., 1996a; Nakayama et al., 1996b; Booton et al., 1998b; Krienitz et al., 2003; Lewis & McCourt, 2004; Watanabe et al., 2006a; Watanabe et al., 2006b; Nakada et al., 2008a; Nakada et al., 2008b). As presently conceived, the class includes non-motile or motile unicells with two or four flagella, biflagellate colonies, filaments and cells that are imbedded in a mucilage envelope (Nakazawa et al., 2004; Nozaki et al., 2006b; Yamada et al., 2008; Nakada & Nozaki, 2009; Matsuzaki et al., 2010; Novis et al., 2010; Nozaki et al., 2010). The Chlamydomonadales mainly comprise freshwater and terrestrial green algae, including several psychrophilic and/or halotolerant species (e.g., Dunaliella and Chlamydomonas nivalis) (Eddie et al., 2008; Buchheim et al., 2010; Muramoto et al., 2010; Remias et al., 2010). A few species have evolved an obligate heterotrophic life style (e.g., Polytoma, Polytomella) and have plastids with highly modified functions and genomes (Vernon et al., 2001). 18S phylogenetic analyses revealed 21 main clades with a poorly resolved backbone (Nakada et al., 2008a). Several genera are polyphyletic, with the most extreme example being the large genus Chlamydomonas, which is distributed in at least five distinct lineages within the Chlamydomonadales (Buchheim et al., 1990; Buchheim et al., 1996; Pröschold et al., 2001; Nakada et al., 2008a). Phylogenetic analyses based on chloroplast genes and 18S sequences showed that quadriflagellate members with CCW or CW basal body orientation (Golenkinia, Carteria, Pseudocarteria, Hafniomonas, Trochiscia, Treubaria and Cylindrocapsa) form early diverging lineages, suggesting that the CW orientation of most chlamydomonadalean genera may have evolved from the CCW orientation in ancestral quadriflagellate members (Nozaki et al., 2003; Nakada et al., 2008a). [update: The early diverging position of the quadriflagellate Tetraflagellochloris further supports this hypothesis (Barsanti et al. 2013)].
Because the Chlamydomonadales include unicellular flagellates (e.g., Chlamydomonas) and colonial forms (e.g., Volvox), its members have been extensively studied as models of multicellular evolution (Kirk, 2003; Kirk, 2005; Herron & Michod, 2008; Herron et al., 2009; Prochnik et al., 2010; Ueki et al., 2010). Interestingly, several phylogenetic studies have demonstrated non-monophyly of colonial flagellates and the absence of a phylogenetic trend in the level of colony complexity, complicating evolutionary interpretations (Buchheim & Chapman, 1991; Buchheim et al., 1994; Herron & Michod, 2008; Nakada et al., 2008a; Nakada et al., 2010). We will return to the subject of evolution of multicellularity in the section “Genomic insights into the evolution of complexity in volvocine green algae”.
The Sphaeropleales form another large group of Chlorophyceae, including some of the most common freshwater phytoplankters such as Scenedesmus, Desmodesmus and Pediastrum, as well as picoplanktonic members (Wolf et al., 2002; Krienitz et al., 2011). Representatives are non-motile unicells, colonies or filaments, which produce biflagellate zoospores (zoosporic) or non-motile spores (autosporic). The filamentous Microspora has been allied with the coccoid genus Bracteacoccus based on ultrastructure of the flagellar apparatus and 18S data (Lokhorst & Star, 1999; Buchheim & Buchheim, 2001). Monophyly of the Sphaeropleales has been supported by unique rDNA ITS2 secondary structure characteristics (Keller et al., 2008) and phylogenetic analysis of nuclear rDNA and plastid atpB and rbcL sequences (Verghese, 2007). Diversity and phylogenetic relationships within the Sphaeropleales have been studied by Lewis (1997), Krienitz et al. (Krienitz et al., 2001; 2003; 2011), Buchheim et al. (2001; 2005), Shoup and Lewis (2003), Wolf et al. (2003), McManus and Lewis (2005; 2011) and Johnson et al. (2007).
In contrast to the species-rich CS clade, the OCC orders are much less diverse. The Chaetophorales of about 10 genera include unbranched (e.g., Uronema) or branched (e.g., Chaetophora, Stigeoclonium) filaments that produce quadriflagellate zoospores. Phylogenetic relationships within the order have been studied based on 18S sequence data (Caisová et al., 2011). The Chaetopeltidales, originally erected based on ultrastructural data (O’Kelly et al., 1994), includes vegetative disk-like thalli (e.g., Chaetopeltis, Pseudulvella) (Sanchez-Puerta et al., 2006), or three-dimensional packets of cells (e.g., Floydiella) that reproduce asexually by quadriflagellate spores. 18S phylogenetic analyses demonstrated that Planophila, which was originally included in the order, is polyphyletic and distributed among the three core ulvophycean classes (Friedl & O’Kelly, 2002). The Oedogoniales includes the filamentous genera Oedogonium, Oedocladium and Bulbochaete. They share a unique form of cytokinesis and a specialized form of oogamous sexual reproduction involving the production of stephanokont motile cells (Pickett-Heaps, 1975). Phylogenetic relationships within the order have been studied by Alberghina et al. (2006) and Mei et al. (2007).
3.4. Ulvophyceae
The Ulvophyceae was originally defined based on a suite of ultrastructural characteristics, including a CCW orientation of the flagellar root system, cytokinesis by furrowing, a closed persistent spindle and the absence of a phycoplast (Mattox & Stewart, 1984; O’Kelly & Floyd, 1984b; O’Kelly & Floyd, 1984a; Sluiman, 1989). The class is best known for its macroscopic marine representatives (the green seaweeds), but several members also occur in freshwater or damp subaerial habitats such as soil, rocks, tree bark and leaves (Chihara et al., 1986; López-Bautista & Chapman, 2003; Rindi et al., 2006; Watanabe & Nakayama, 2007).
In terms of diversity in thallus complexity and cellular sophistication, the Ulvophyceae far exceed the other chlorophytan classes. Their morphologies range from microscopic unicells to macroscopic multicellular plants, and giant-celled organisms with unique cellular and physiological characteristics (Mine et al., 2008). Four main cytomorphological types can be distinguished (Cocquyt et al., 2010b). The first type comprises non-motile uninucleate unicells, and is present in some Ulotrichales (Chihara et al., 1986; Nakayama et al., 1996a; Friedl & O’Kelly, 2002; Watanabe & Nakayama, 2007) and a number of genera of uncertain affinity (e.g., Oltmannsiellopsis and Ignatius). The second type consists of multicellular filaments or blades composed of uninucleate cells; it characterizes most Ulvales, Ulotrichales and Trentepohliales. The third type has multicellular bodies composed of multinucleate cells with nuclei organized in regularly spaced cytoplasmic domains (McNaughton & Goff, 1990; Motomura, 1996). This is known as the siphonocladous type and characterizes the Cladophorales and Blastophysa and some members of the Ulotrichales (e.g., Urospora and Acrosiphonia). The fourth type is better known as siphonous type and is characterized by plants consisting of a single giant tubular cell. It is present in the orders Bryopsidales and Dasycladales. Siphonous cells generally contain thousands to millions of nuclei. Many Dasycladales, however, remain uninucleate throughout much of their life cycle with a giant diploid nucleus that only divides at the onset of reproduction (Berger & Kaever, 1992). In contrast to the siphonocladous type, the cytoplasm of siphonous algae exhibits vigorous streaming, enabling transportation of transcripts across the plant (Menzel, 1987; Menzel, 1994; Mine et al., 2001). Although some siphonous algae are tiny microscopic siphons, many form large and complex seaweeds that exhibit morphological differentiation into structures that resemble the roots, stems, and leaves of land plants and even have similar functions (Chisholm et al., 1996). The evolution of siphonocladous and siphonous architectures coincided with several cytological and cytoskeletal specializations such as unique mechanisms of wounding response (Menzel, 1988; La Claire, 1992; Kim et al., 2001; Mine et al., 2008). In addition to facilitating transport of transcripts as mentioned above, the evolution of cytoplasmic streaming in siphonous algae also allowed transport of nutrients and organelles throughout the siphonous algal body. In combination with morphological changes, this allows nutrient uptake from marine sediments (Chisholm et al., 1996) and chloroplast migration to optimize photosynthesis and avoid herbivory by micrograzers. Such innovations have most likely had selective advantages and contribute to the ecological dominance of siphonous algae in tropical and warm-temperate coastal ecosystems (Vroom & Smith, 2001).
Understanding the phylogenetic relationships among the ulvophycean orders can provide important insights into the diversification of cytological types and evolution of morphological complexity in the class. Analyses of 18S datasets have supported the circumscription of traditional orders based on cytomorphological characteristics but could not resolve the relationships among them (Chappell et al., 1991; López-Bautista & Chapman, 2003; Watanabe & Nakayama, 2007; Leliaert et al., 2009a). Moreover, the monophyly of the class has been questioned because it lacks unique ultrastructural synapomorphies (Mattox & Stewart, 1984; O’Kelly & Floyd, 1984a). Several molecular phylogenetic studies provided weak or no support for monophyly of the Ulvophyceae and recovered two distinct lineages: the Oltmannsiellopsidales–Ulvales–Ulotrichales lineage and a clade consisting of Trentepohliales and the siphonocladous and siphonous seaweed orders (Zechman et al., 1990; Watanabe et al., 2001; López-Bautista & Chapman, 2003; Watanabe & Nakayama, 2007; Cocquyt et al., 2009; Zuccarello et al., 2009). More recently, a 10-gene phylogenetic analysis recovered the class as a well supported monophyletic group, and confirmed the divergence of two main ulvophycean clades (Cocquyt et al., 2010b). These two discrete clades are also supported by independent molecular data, including the distribution of elongation factor genes and a deviant genetic code (Cocquyt et al., 2009; Gile et al., 2009; Cocquyt et al., 2010a). The phylogeny of Cocquyt et al. (2010b) recovered the Trentepohliales as the sister lineage of the Bryopsidales, Cladophorales, and Dasycladales. The Bryopsidales and Dasycladales were recovered as sister lineages, and the genus Blastophysa was sister to the Cladophorales. The relationship of the enigmatic endophytic green alga Blastophysa with the Cladophorales is corroborated by morphological, ultrastructural, cytological, and biochemical features (O’Kelly & Floyd, 1984a; Chappell et al., 1991). On the other hand, the grouping of the Dasycladales and Bryopsidales was surprising given that ultrastructural features of the flagellar apparatus suggested that Dasycladales may be more closely related to Cladophorales (O’Kelly & Floyd, 1984a).
The phylogeny of Cocquyt et al. (2010b) has provided the groundwork for new insights into the cytomorphological diversification of the Ulvophyceae from an ancestral uninucleate unicell. The phylogeny indicates that macroscopic growth was achieved independently in various lineages involving radically different mechanisms: either by evolving multicellularity with coupled mitosis and cytokinesis (Ulvales-Ulotrichales and Trentepohliales), or by obtaining siphonocladous or siphonous organizations. The evolution of siphonous and siphonocladous cytologies is hypothesized to have evolved independently from a unicellular ancestor as a result of selective pressures for macroscopic growth in marine benthic environments. In this view, two pathways toward enlarged cells and macroscopic growth would have emerged. The first involved the evolution of multinucleate cells where every nucleus provides for its cytoplasmic domain, leading to the siphonocladous cytological organisation of the Cladophorales and Blastophysa. The second pathway, which led to the siphonous Bryopsidales and Dasycladales, involved two steps. Initially, the enlarged ancestral cell developed a macronucleus and cytoplasmic streaming, allowing increased transcription from a single nucleus and distribution of transcripts across the cell. The second step involved the evolution of multinucleate siphons, which possibly occurred independently in both orders in association with the evolution of complex, macroscopic plants that have outgrown the potential of a single macronucleus.
The Ulvales-Ulotrichales forms a diverse clade of unicells and multicellular thalli, ranging from branched or unbranched filaments to blades or tubular forms. The group is predominantly marine, but several transitions to freshwater or terrestrial habitats have occurred independently (Shimada et al., 2008; Ichihara et al., 2009; Suutari et al., 2010). The orders Ulvales and Ulotrichales have traditionally been distinguished based on life history features: the Ulvales have a life cycle involving isomorphic alternations of multicellular stages, while in the Ulotrichales the diploid, spore-producing stage is a small, thick-walled unicell that is attached to the substrate by a stalk (Codiolum stage) (O’Kelly et al., 2004b). Molecular phylogenetic studies provide poor or no support for the separation of the orders (e.g., O’Kelly et al., 2004a; O’Kelly et al., 2004c). Diversity and phylogenetic relationships within the clade have been studied based on plastid genes (mainly rbcL) and nuclear 18S sequence data (Hayden & Waaland, 2002; Hayden & Waaland, 2004; O’Kelly et al., 2004a; O’Kelly et al., 2004c; O’Kelly et al., 2004b; Lindstrom & Hanic, 2005; Lindstrom et al., 2006; Loughnane et al., 2008; Heesch et al., 2009; Kraft et al., 2010; O’Kelly et al., 2010). The relationships of unicellular or sarcinoid members (e.g., Desmochloris, Halochlorococcum, Pseudoneochloris) may provide important insights into the evolution of multicellularity in the clade, but as yet, their phylogenetic positions are poorly resolved (Watanabe et al., 2001; O’Kelly et al., 2004a; O’Kelly et al., 2004c).
The Oltmannsiellopsidales is generally inferred to have diverged near the base of the Ulvales-Ulotrichales clade (Watanabe & Nakayama, 2007; Cocquyt et al., 2009; Leliaert et al., 2009a). The order consists of a small number of marine and freshwater species. Oltmannsiellopsis includes quadriflagellate unicells or small colonies (Nakayama et al., 1996a). Dangemannia microcystis and Halochlorococcum marinum are non-motile unicells, disks or packets of cells that produce quadriflagellate spores (Friedl & O’Kelly, 2002; Pröschold et al., 2002).
[Update: Scotinosphaerales forms a distinct and highly divergent clade of Ulvophyceae (Škaloud et al. 2013). The order consist of a single genus, Scotinosphaera, from freshwater or damp terrestrial habitats, which is characterized by large cells with several unique ultrastructural features (Škaloud et al. 2013). The early diverging position of the Scotinophaerales enforces the notion that nonmotile unicellular freshwater organisms have played an important role in the early diversification of the Ulvophyceae.]
The Cladophorales (including Siphonocladales) comprises four main lineages, likely from a filamentous marine ancestor (Bakker et al., 1994; Hanyuda et al., 2002; Leliaert et al., 2003; Leliaert et al., 2009a). The marine unbranched, microfilamentous Okellya forms the earliest diverging lineage, followed by the Aegagropila clade, which is often found in highly specialized freshwater habitats (e.g., lake-balls or epizoophytes on turtles and snails). The species-rich Cladophora clade, including branched (Cladophora) or unbranched (Chaetomorpha and Rhizoclonium) filaments, is predominantly marine, but at least one clade has invaded freshwater habitats. The marine tropical Siphonocladus clade is morphologically most diverse, comprising highly specialized forms and giant-cells with unique cytomorphological traits and modes of cell division (e.g., Valonia, Boergesenia, Dictyosphaeria) (Okuda et al., 1997a; Okuda et al., 1997b; Mine et al., 2008). Molecular phylogenies have shown that these specialized features have evolved independently multiple times in the clade (Leliaert et al., 2007). The large genus Cladophora, and several other genera have been found to be polyphyletic (Leliaert et al., 2003; Leliaert et al., 2009b).
The Bryopsidales (also referred to as the Caulerpales, Codiales or Siphonales) range in morphology from simple, branched siphons (e.g., Bryopsis, Chlorodesmis) to more complex, differentiated thalli (e.g., Codium, Halimeda, Udotea, Caulerpa). Several species form key components of tropical marine coastal ecosystems, where they are among the major primary producers on coral reefs, in lagoons and seagrass beds. The thallus surface of several species is calcified, and some are important contributors to coral reef structure. A single genus, Dichotomosiphon, occurs in freshwater habitats. A phylogeny based on rbcL sequence data (Lam & Zechman, 2006) revealed two main clades, corresponding to the suborders Bryopsidineae and Halimedineae, which are characterized based on differences in thallus morphology, reproduction and plastid types (Hillis-Colinvaux, 1984). Increased gene and taxon sampling improved phylogenetic resolution in these clades and revealed the limestone-boring genus Ostreobium as an early diverging lineage (Verbruggen et al., 2009a). Furthermore, non-monophyly has been demonstrated for several genera and species (Verbruggen et al., 2009b).
The Dasycladales are marine tropical algae characterized by radially symmetrical thalli encrusted with calcium carbonate. Two families have traditionally been recognized, Dasycladaceae and Polyphysaceae, based on differences in the reproductive structures. Early phylogenetic studies based on 18S or rbcL sequence data indicated that at least the Dasycladaceae was not monophyletic (Olsen et al., 1994; Berger et al., 2003; Zechman, 2003). A 5-gene phylogeny with extended taxon sampling demonstrated that the Dasycladaceae form a paraphyletic assemblage that gave rise to a monophyletic Polyphysaceae (Verbruggen et al., 2009a). The Dasycladales have a rich fossil record and were much more diverse historically than they are today.
The Trentepohliales is an entirely terrestrial order and was originally omitted from the Ulvophyceae based on atypical characteristics such as a multilayered structure (MLS) in the flagellar root system, phragmoplast-like cytokinesis, the presence of plasmodesmata between vegetative cells, and a unique type of sporangial reproduction implying a relationship with streptophyte green algae (Chapman, 1984; O’Kelly & Floyd, 1984a; Chapman et al., 2001). However, molecular data firmly established an alliance with the ulvophycean orders Cladophorales, Bryopsidales and Dasycladales (Zechman et al., 1990; López-Bautista & Chapman, 2003; Cocquyt et al., 2010b), either pointing towards parallel evolution of the streptophyte-like ultrastructural features or indicating that some of these characteristics (e.g., a MLS) may represent an ancestral condition in the green lineage (Lewis & McCourt, 2004); either way it is highly interesting that a phragmoplast-like cytokinesis has evolved in Trentepohliales and land plants, both found in terrestrial environments. The phylogenetic position of the Trentepohliales within a clade of marine orders suggests a sea-to-land transition, which would be unique among algae (Lewis & McCourt, 2004). Diversity and phylogenetic relationships in the order have been studied based on rbcL and 18S rDNA sequence data (López-Bautista et al., 2006; Rindi et al., 2009).
4. Streptophyta: charophyte green algae and the origin of land plants
As one of the deepest branches of the green lineage, the Streptophyta are certainly old, quite likely a billion years or more, and both molecular sequence divergence and the few available fossils are consistent with that view. However, despite this great age, most of the subclades of Streptophyta are not particularly diverse, with the striking exception of the embryophytes (land plants), a lineage with several hundred thousand species.
Ultrastructural, biochemical and molecular data revealed six distinct groups of charophytes: Charophyceae, Coleochaetophyceae, Zygnematophyceae, Klebsormidiophyceae, Chlorokybophyceae and probably the Mesostigmatophyceae (Mattox & Stewart, 1984; Qiu & Palmer, 1999; McCourt et al., 2004). Considerable progress has been made during the past decade in clarifying the relationships among these lineages, and elucidating the closest living relative of the land plants (Karol et al., 2001; Turmel et al., 2003; Qiu et al., 2006; Turmel et al., 2006b; Lemieux et al., 2007; Rodríguez-Ezpeleta et al., 2007; Finet et al., 2010; Wodniok et al., 2011).
4.1. The early diverging charophytes
Mesostigma and Chlorokybus likely form the earliest diverging lineage of extant streptophytes (Bhattacharya et al., 1998; Simon et al., 2006; Turmel et al., 2006a; Lemieux et al., 2007; Rodríguez-Ezpeleta et al., 2007), although some early chloroplast and mitochondrial phylogenomic analyses with more limited taxon sampling suggested that Mesostigma emerged before the divergence of the Streptophyta and Chlorophyta (Lemieux et al., 2000; Turmel et al., 2002c). Additional evidence for the placement of Mesostigma in the Streptophyta comes from the presence of a MLS in the flagellar root system (Rogers et al., 1981; Melkonian, 1989), a shared multigene family (BIP) between Mesostigma and the rest of the Streptophyta (Nedelcu et al., 2006), and a shared duplication of the GapA/GapB gene (Petersen et al., 2006). Plastid phylogenomic analyses implied a sister relationship between Mesostigma and Chlorokybus (Lemieux et al., 2007; Rodríguez-Ezpeleta et al., 2007), while two nuclear multi-gene phylogenies suggested that Chlorokybus diverged after Mesostigma (Cocquyt et al., 2010b; Finet et al., 2010). The non-sister relationship between Mesostigma and Chlorokybus is further supported by the fact that Mesostigma is the only streptophyte with flagella in its vegetative stage (a presumed ancestral feature of the green plant lineage). This suggests that loss of flagella occurred after the divergence of Mesostigma (Fig. 4).
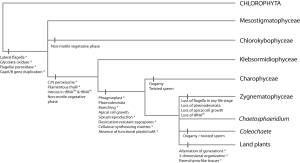
Morphologically, Mesostigma and Chlorokybus are among the simplest streptophyte algae. Mesostigma (the only genus in the Mesostigmatophyceae) is a freshwater, scaly, asymmetrical unicell with two flagella (Marin & Melkonian, 1999) and a unique suite of photosynthetic pigments (Yoshii et al., 2003; Hall & Delwiche, 2007). Chlorokybus (the sole member of the Chlorokybophyceae) occurs in moist terrestrial habitats where it forms packets of a few cells that reproduce asexually by asymmetrical, motile spores with ultrastructural streptophyte characteristics (Rogers et al., 1980; Lokhorst et al., 1988). Sexual reproduction is unknown in the two lineages. Spirotaenia, which is normally placed in the Zygnematophyceae based on the lack of flagella and sexual reproduction by conjugation, was unexpectedly inferred as a sister lineage to Chlorokybus based on 18S rDNA and chloroplast rbcL sequence data (Gontcharov & Melkonian, 2004).
Multi-gene phylogenies show that the Klebsormidiophyceae diverged after the Mesostigmatophyceae and Chlorokybophyceae, and is sister to the clade comprising the Charophyceae, Zygnematophyceae, Coleochaetophyceae and land plants (Karol et al., 2001; Turmel et al., 2002a; Cocquyt et al., 2010b; Finet et al., 2010; Wodniok et al., 2011). This phylogenetic position is further supported by some chloroplast genomic features, like the presence of introns in two transfer RNAs which is shared between Klebsormidium and the later-diverging streptophyte lineages, and the presence of functional plastid tufA, which is shared with the early-diverging lineages (Baldauf et al., 1990; Baldauf & Palmer, 1990; Manhart & Palmer, 1990; Turmel et al., 2005; Turmel et al., 2007b) (Fig. 4). The Klebsormidiophyceae includes the freshwater or terrestrial genera Klebsormidium, Interfilum and Entransia, which form unbranched filaments or sarcinoid packets that produce asexual motile spores with streptophyte characteristics (Cook, 2004b; Sluiman et al., 2008). Diversity and phylogenetic relationships within the Klebsormidiophyceae have been studied by Rindi et al. (2008; 2011) and Mikhailyuk et al. (2008).
In contrast to the three early-diverging streptophyte lineages (Mesostigmatophyceae, Chlorokybophyceae and Klebsormidiophyceae), which undergo cytokinesis by furrowing, the remaining lineages produce a phragmoplast and plasmodesmata (absent in the Zygnematophyceae), facilitating intercellular communication and differentiation (Pickett-Heaps, 1975; Raven, 1997). Additional traits that are uniquely shared by the remaining streptophytes include apical growth (not in the Zygnematophyceae), sexual reproduction, some biochemical features, and similar cellulose-synthesizing rosettes (Tsekos, 1999).
4.2. The closest living relative of the land plants
Numerous studies have focused on identifying the closest living relative to the land plants, and several lineages have been proposed based on morphological, ultrastructural and molecular data (Graham, 1984; Graham, 1993; Kranz et al., 1995; Bhattacharya & Medlin, 1998; An et al., 1999; Graham et al., 2000; McCourt et al., 2004). Multi-marker phylogenies have been sensitive to taxon and gene sampling and provided equivocal results, suggesting the Charophyceae (Karol et al., 2001; Turmel et al., 2007a; Cocquyt et al., 2010b), Zygnematophyceae (Turmel et al., 2006a; Lemieux et al., 2007; Rodríguez-Ezpeleta et al., 2007; Wodniok et al., 2011), Coleochaetophyceae (Turmel et al., 2009a; Turmel et al., 2009b; Finet et al., 2010) as the sister lineage of the embryophytes. Interestingly, mitochondrial and plastid data have yielded inconsistent positions for the Charales (Turmel et al., 2006a; Turmel et al., 2007a), which might result from lateral transfer of large genomic segments or even whole organellar genomes (Turmel et al., 2008).
Two recent multi-gene phylogenetic analyses based on nuclear-encoded genes either placed the Coleochaetophyceae or Zygnematophyceae as sister to the land plants (Finet et al., 2010; Wodniok et al., 2011).
A first analysis based on 77 nuclear ribosomal protein genes suggested that Coleochaete represents the closest relative of land plants (Finet et al., 2010), which agrees with earlier morphology-based hypotheses (Graham, 1984; Graham, 1993; Graham et al., 2000). Coleochaete ranges in morphology from filamentous to relatively complex discoid parenchymatous thalli (Graham, 1982; Graham, 1984; Dupuy et al., 2010). In some species the zygote is retained on the maternal plant and corticated after fertilization by a layer of sterile cells, and receives nourishment via placental transfer cells with wall ingrowths. The walls of the zygote possess sporopollenin, a highly resistant substance found in the outer wall of pollen (Delwiche et al., 1989). Cytokinesis and phragmoplast formation in Coleochaete are typical of land plants (Marchant & Pickett-Heaps, 1973; Graham et al., 2000; Cook, 2004a). However, there is also reason to be cautious. First, an unexpected result in the phylogeny of Finet et al. (2010) was the grouping of Chaetosphaeridium with the Zygnematophyceae. Chaetosphaeridium has typically been placed in the Coleochaetophyceae, a position that has been supported by molecular phylogenetic studies (Karol et al., 2001; Turmel et al., 2002a), and morpho-cytological characteristics, such as the presence of typical sheathed hairs and similar chloroplast structure (Delwiche et al., 2002; Hall & Delwiche, 2007). It is certainly possible that the two genera, both of which live on aquatic plants or stones in freshwater environments, do not form a monophyletic group, but they share enough distinctive properties to cast doubt on a phylogeny that separates them. Second, only limited taxon sampling is currently available for multi-gene analyses, and the deep divergence times involved make dense taxon sampling extremely important to avert systematic error in phylogenetic analyses. [Update: Re-analysis of the dataset of Finet et al. (2010), excluding contaminant sequences, showed that land plants are sister to a group composed of Zygnematophyceae and Coleochaetophyceae (Laurin-Lemay et al. 2012, Finet et al. 2012). In addition, Coleochaetophyceae were recovered monophyletic (see updated Figs 2 and 3)].
A second analysis, based on 129 nuclear genes and slightly fewer taxa, suggested the Zygnematophyceae or a clade consisting of the Zygnematophyceae and Coleochaetophyceae as sister lineage of the land plants (Wodniok et al., 2011). The sister relation between Zygnematophyceae and land plants is supported by some shared features, including some components of the auxin signalling machinery (De Smet et al., 2011) and chloroplast movement (Wada et al., 2003). It is more difficult to find morphological synapomorphies for the relationship between Zygnematophyceae and embryophytes, possibly as a result of secondary simplification and specialization in the Zygnematophyceae (Wodniok et al., 2011). [Update: A sister relationship between Zygnematophyceae and land plants was also supperted by a 160-nuclear-gene phylogenomic analysis of Timme et al. (2012)]
The Zygnematophyceae, also known as conjugating green algae, is the most species-rich and morphologically diverse lineage of charophytes. Vegetative bodies include non-motile unicells, filaments and small colonial forms. Sexual reproduction occurs by a unique process of conjugation, involving fusion of non-motile gametes. Absence of flagellate reproductive stages and basal bodies also sets this class apart from other charophytes. Traditionally, the class was divided into the Zygnematales and the Desmidiales based primarily on differences in cell wall structure, but molecular phylogeneties have shown that the Zygnematales (characterized by smooth non-ornamented cell walls) is a paraphyletic assemblage that gave rise to the monophyletic Desmidiales (McCourt et al., 2000; Gontcharov et al., 2003; Gontcharov et al., 2004; Gontcharov, 2008). The latter order consists of unicells, filaments and colonies with conspicuously ornamented cell walls and cells constricted in two half cells (semicells). The diversity and phylogenetic relationships within the lineage have been studied with nuclear rDNA and chloroplast rbcL sequence data (Denboh et al., 2001; Drummond et al., 2005; Gontcharov & Melkonian, 2005; Kim et al., 2006; Gontcharov & Melkonian, 2008; Hall et al., 2008a; Hall et al., 2008b; Gontcharov & Melkonian, 2010; Gontcharov & Melkonian, 2011).
The Charophyceae (also known as stoneworts) are large and morphologically complex macroscopic algae with a plant-like appearance, consisting of a central stalk of large, elongate and multinucleate cells and whorls of branches radiating from uninucleate cells at the nodes. Growth is by a single apical meristematic cell. Sexual reproduction is oogamous with motile sperm produced in complex antheridia. Oogonia and antheridia are surrounded by sterile cells, and zygotes develop a thick covering of sporopollenin. Because the sporopollenin surrounding the zygotes is highly resistant to degradation and because the shallow freshwater habitats in which they live often create favourable depositional conditions, the stoneworts are well represented in the fossil record (fossilized zygote walls are called gyrogonites). Unambiguous fossils of stoneworts date back to at least 380 mya (Feist & Feist, 1997), with abundant and diverse gyrogonites in more recent sediments. Phylogenetic relationships within the class have been studied by McCourt et al. (1996; 1999) Sakayama et al. (2004; 2005; 2008; 2009).
As noted above, Karol et al. (2001) presented a four-gene phylogeny with relatively dense taxon sampling that placed the Charophyceae as the sister taxon to embryophytes. Thus, plausible recent analyses have shown almost every possible arrangement among the four lineages of “higher” streptophytes or phragmoplastophytina (embryophytes, Coleochaetophyceae, Zygnematophyceae and Charophyceae). Given the very deep divergence times involved, it is not surprising that representation by six or eight species is not sufficient to provide a well-resolved phylogeny even when multiple genes are available for analysis. What is far less clear is whether or not enough diversity has survived extinction to permit taxon sampling alone to resolve this phylogenetic puzzle, or whether new analytical approaches will be needed to distinguish among the alternative topologies.
We will return to the subject of land plant origins in the section “Molecular evolution in the Streptophyta and the origin of land plants”, in which we discuss molecular evolution in the streptophyte lineage, emphasizing the genetic facilitation of land plant origins.
III. Spread of green genes in other eukaryotes
A growing body of data is providing evidence for widespread movement of genetic information between distantly related eukaryotic genomes (Keeling & Palmer, 2008; Schaack et al., 2010). Horizontal gene transfer (HGT, also known as lateral gene transfer) is a mechanism to rapidly spread evolutionary innovations across distinct lineages and is now regarded as an important force in the evolution of eukaryotes and their genomes (Nedelcu et al., 2008; Sun et al., 2010). Green algal genes have been incorporated into the genomes of various unrelated eukaryotes via different mechanisms (Fig. 2).
Two important modes of eukaryote-to-eukaryote HGT are the gene ratchet mechanism and endosymbiosis (Dagan & Martin, 2009a; Sun et al., 2010). The gene ratchet mechanism, also known as “you are what you eat”, posits that by engulfing and digesting other cells, phagotrophic protists may occasionally incorporate genetic material of their prey into their genomes (Doolittle, 1998). In a few instances during eukaryote evolution, captured cells have been retained as intracellular symbionts (endosymbionts) and have become stably integrated inside the host cell. The two most prominent endosymbioses in eukaryote evolution have involved an alpha-proteobacterial and a cyanobacterial endosymbiont, and have given rise to mitochondria and plastids, respectively (Keeling & Palmer, 2008). Both endosymbionts have undergone massive gene loss and gene transfer from the endosymbiont genome to the host genome (endosymbiotic gene transfer, EGT), reducing them to metabolic slaves (organelles) inside their host cells (Gould et al., 2008; Tirichine & Bowler, 2011). The plastids of the green lineage, the red algae and glaucophytes are derived from a single primary endosymbiotic event, i.e. the original uptake and retention of a cyanobacterium by a nonphotosynthetic eukaryotic host cell (Rodríguez-Ezpeleta et al., 2005). The great majority of other photosynthetic eukaryotes acquired their plastids by secondary or tertiary endosymbioses, in which eukaryotes with green or red plastids were preyed upon by a heterotrophic eukaryote, followed by EGT, this time from the endosymbiont nucleus to that of the secondary or tertiary hosts (Delwiche, 1999; Keeling, 2004; Rodríguez-Ezpeleta et al., 2005; Keeling, 2010). The number of secondary (and tertiary) endosymbiotic events is heavily debated (Archibald, 2009).
Three eukaryotic lineages have fully integrated secondary plastids of green algal origin: photosynthetic euglenids, chlorarachniophytes and the “green” dinoflagellates (Delwiche, 1999; Keeling, 2004; Archibald, 2009; Keeling, 2010). While it has been proposed that the chloroplasts of chlorarachniophytes and euglenids share a common origin (the Cabozoa hypothesis, Cavalier-Smith, 1999), chloroplast multi-gene phylogenies suggest that these plastids originated independently (Rogers et al., 2007; Takahashi et al., 2007; Turmel et al., 2009a). The Pyramimonadales have been identified as the source of the chloroplasts of the euglenid Euglena gracilis (Turmel et al., 2009a; Matsumoto et al., 2011) (Fig. 2). The chloroplast of the chlorarachniophyte Bigelowiella natans was found to be sister to the ulvophytes Pseudendoclonium and Oltmannsiellopsis (Turmel et al., 2009a), although this relationship was not supported in an analysis based on more taxa but fewer genes (Matsumoto et al., 2011).
Euglenids are a diverse group of mainly freshwater flagellate unicells, including photosynthetic members with plastids bounded by three membranes, as well as nonphotosynthetic members (Keeling, 2004; Triemer & Farmer, 2007). Together with the kinetoplastids and diplonemids they make up the Euglenozoa, which belong to the Excavata, a largely nonphotosynthetic eukaryotic supergroup (Baldauf, 2008). Chloroplast genomes have been characterized for two euglenids: the photosynthetic E. gracilis and the closely related, nonphotosynthetic E. longa (Hallick et al., 1993; Gockel & Hachtel, 2000). The chloroplast genome of E. gracilis is 143 kbp in size and shows high similarities in gene order with Pyramimonas (Turmel et al., 2009a). The plastid genome of Euglena longa is highly reduced (73 kbp) and has lost most photosynthesis-related genes but retained most of the other genes present in E. gracilis.
Chlorarachniophytes are a small group of marine amoeboflagellates bearing plastids bounded by four membranes (Keeling, 2004; Ishida et al., 2007). The discrepancy of chloroplasts with three membranes in euglenids versus four membranes in chlorarachniophytes has been suggested to have resulted from a myzocytosic aquisation of the plastids in euglenids versus a phagocytosic capture in chlorarachniophytes (Keeling, 2010). Chlorarachniophytes belong to the Rhizaria, which, like the Excavata, are primarily nonphotosynthetic (Baldauf, 2008). Chlorarachniophytes are one of only two groups of photosynthetic eukaryotes (the other one being the red plastid-containing cryptophytes) in which remnant nuclei of the eukaryotic endosymbionts, called nucleomorphs, have been retained (Cavalier-Smith, 2002). Chlorarachniophyte nucleomorphs contain highly reduced genomes (330-610 Kbp) and most essential genes have been transferred to the host’s nuclear genome. The nucleomorphs contain three small linear chromosomes and a gene density similar to that seen in prokaryotes (Moore & Archibald, 2009). Remarkably, the nucleomorph genomes of chlorarachniophytes and cryptophytes have evolved similar genomic features, including a three-chromosome architecture with subtelomeric rRNA operons. The 69 Kbp chloroplast genome of Bigelowiella natans is smaller than plastids of most other green algae and is closer in size to genomes of several nonphotosynthetic plastids (Rogers et al., 2007). Unlike nonphotosynthetic plastids, however, the B. natans chloroplast genome is highly compacted and encodes most of the genes found in other photosynthetic green algal plastids.
Dinoflagellates are a diverse group of photosynthetic or heterotrophic flagellates. While most photosynthetic dinoflagellates have secondary or tertiary plastids of red algal origin, members of the “green” dinoflagellate genus Lepidodinium have replaced their original red plastids with a new secondary endosymbiont from the green lineage (termed serial secondary endosymbiosis) (Watanabe et al., 1990; Keeling, 2010). Although prasinophytes had been hypothesized as the chloroplast source of Lepidodinium based on pigment composition (Watanabe et al., 1990), recent chloroplast multi-gene phylogenetic analyses suggest that these plastids were derived from an early representative of the core chlorophytes, but the exact donor lineage remains equivocal (Takishita et al., 2008; Matsumoto et al., 2011).
In addition to these stable endosymbioses, there is also evidence for early stages of plastid acquisition via secondary endosymbioses (Gould et al., 2008). The katablepharid flagellate Hatena arenicola harbours a prasinophyte green algal endosymbiont of the genus Nephroselmis (Okamoto & Inouye, 2005; Okamoto & Inouye, 2006). The single membrane-bound endosymbiont exhibits extensive structural changes when within the Hatena cell and is ultrastructurally tightly associated with its host. However, the integration of the symbionts is not entirely stable as the division of the symbiont is not coordinated with that of the host: cell division results in only one symbiont-bearing daughter cell while the other daughter cell re-establishes a phototrophic lifestyle by capturing a new Nephroselmis symbiont from the environment. As yet, it is unknown if this endosymbiotic association has resulted in EGT.
A remarkable case of endosymbiosis is the plastid theft performed by sacoglossan molluscs (Elysia and relatives) (Gould et al., 2008). These sea slugs feed upon siphonous and siphonocladous green algae (Ulvophyceae) and siphonous xanthophytes (e.g., Vaucheria). Some species are able to sequester chloroplasts from these algae into their gut cells (Händeler et al., 2009; Händeler et al., 2010). Despite the absence of algal nuclei, these so-called kleptoplasts remain transcriptionally and translationally active and allow the slugs to rely on photosynthate for weeks to months (Mujer et al., 1996). The kleptoplasts are not permanently acquired; they are unable to divide inside the host and are not passed on from one slug generation to another. The molecular basis of plastid retention is not well understood but has been suggested to involve gene transfer from the algal food source to the slug’s genome. Indeed, recent studies have provided evidence of gene transfer in a species retaining xanthophyte plastids (Rumpho et al., 2008; Schwartz et al., 2010). Conversely, analysis of transcriptome data from two other species maintaining green algal plastids did not reveal any transfer of algal genes specific to photosynthetic function, suggesting that these slugs do not express genes acquired from algal nuclei to maintain plastid function (Wägele et al., 2011).
There are several other examples of green algae that persist as endosymbionts in various eukaryotic hosts. The ciliate Paramecium bursaria engages in close physiological association with trebouxiophyte green algal endosymbionts (Chlorella and relatives) that reside in vacuoles close to the cell surface and are vertically transmitted to daughter cells upon host cell division (Hoshina & Imamura, 2008; Nowack & Melkonian, 2010). This symbiosis has been found to be facultative in lab conditions since both the Paramecium and the algae can be cultivated separately. Other eukaryotes that harbour green algal symbionts (including parasites) include fungi, dinoflagellates, foraminifers, radiolarians, sponges, marine flatworms, cnidarians, molluscs (nudibranchs and giant clams) and vertebrates (Parke & Manton, 1967; Sweeney, 1976; Cachon & Caram, 1979; Williamson, 1979; Friedl & Bhattacharya, 2002; Lewis & Muller-Parker, 2004; Rodriguez et al., 2008; Kovacevic et al., 2010; Nowack & Melkonian, 2010; Kerney et al., 2011). As yet, it remains unclear whether these associations have resulted in endosymbiont-to-host gene transfer.
Remarkably, green algal-derived genes have also been detected in the genomes of a diverse array of other eukaryotes without green algal plastids, including algae with red-type plastids and eukaryote lineages without plastids. Phylogenomic analyses of two diatom genomes revealed that a considerable proportion of nuclear genes are of green algal origin, which is surprising given that diatom plastids are of red algal origin (Moustafa et al., 2009). These findings have been interpreted as evidence for the presence of a green algal endosymbiont early in the evolution of the chromalveolate lineage, which was later replaced by an endosymbiont with a red-type plastid (Moustafa et al., 2009). An ancient green algal endosymbiont may also explain the presence of numerous “green” genes in oomycetes (nonphotosynthetic chromalveolates) and apicomplexan parasites (chromalveolates containing nonphotosynthetic plastids of red-algal origin) (Huang et al., 2004; Tyler et al., 2006; Janouškovec et al., 2010). Similarly, the presence of genes related to green algal sequences in trypanosomatid parasites (kinetoplastids) has been explained by EGT of an ancient green algal endosymbiont (Hannaert et al., 2003), but these data are open to interpretation and scenarios of ancient cryptic secondary endosymbioses in the chromalveolates and other eukaryotes have been questioned (Dagan & Martin, 2009b; Elias & Archibald, 2009; Stiller et al., 2009; Sun et al., 2010). In Monosiga, a member of the Choanozoa, which forms the sister group of the Metazoa, numerous genes of putative algal origin are present, including several genes with green algal affinities (Nedelcu et al., 2008; Sun et al., 2010). Based on the taxonomic distribution pattern of the algal genes and the phagotrophic nature of choanoflagellates, the presence of algal genes in Monosiga has been explained by the gene ratchet mechanism rather than ancient cryptic endosymbiosis (Sun et al., 2010).
IV. Green algal evolution: insights from genes and genomes
The large diversity of green plants offers a unique opportunity to study the molecular mechanisms underlying its evolution. The rapid accumulation of genomic data, along with current phylogenetic hypotheses, is greatly advancing our understanding of molecular evolution in the green lineage. Apart from vertical descent with modification, various other evolutionary events may affect gene histories, including gene duplication, gene loss, horizontal gene transfer and gene rearrangements (Koonin, 2005). Gene duplications and horizontal gene transfer have been suggested to contribute to biological novelty in the green lineage. Large-scale gene and whole genome duplication events have been well-characterized in embryophyte lineages (Flagel & Wendel, 2009) but have also played an important role in the evolution of phenotypic novelty within green algal lineages (Petersen et al., 2006; Robbens et al., 2007b; Becker & Hoef-Emden, 2009). Several studies have indicated horizontal transfers of genetic information from green algae to other eukaryotes or vice versa (Bhattacharya et al., 1996; Friedl et al., 2000; Cocquyt et al., 2009; Ghoshroy et al., 2010).
Below we focus on a number of specific topics dealing with molecular evolution in the green lineage. A first section summarizes our current understanding of chloroplast and mitochondrial genome evolution in green algae. A second section reviews genomic insights into the ecology of oceanic picoplanktonic prasinophytes. Thirdly, molecular mechanisms underlying the evolution of multicellularity and cellular differentiation in volvocine green algae is discussed. A fourth section discusses the evolution of genetic codes and the translational apparatus in green seaweeds. Finally, we discuss recent progress in our understanding of molecular evolution in the streptophyte lineage, focusing on the genetic bases underlying the origin of land plants.
A. Organelle genome evolution
The exploration of organelle genomes — the DNA molecules found within the mitochondria and plastids of eukaryotic cells — has revealed an astonishing array of genomic architectures (Palmer, 1985; Lang et al., 1999). Some of the most diverse and unusual mitochondrial and plastid DNAs (mtDNAs and ptDNAs) from all eukaryotes come from green algae (Gray, 1999). Indeed, among green algal groups there is an impressive range of organelle genome sizes (13 to >525 kb), conformations (circular or linear), chromosome numbers (monomeric, bipartite, or highly fragmented), compactnesses (20-95% coding DNA), gene repertoires (10 to >130 genes), and nucleotide compositions (20 to >60% guanine and cytosine) — see Table 1 for examples and references. Investigations of green algal organelle DNAs have helped unravel the origins and interrelationships of green plants and eukaryotes as a whole (Lemieux et al., 2000; Turmel et al., 2006a; Archibald, 2009), provided insights into the forces driving genome evolution (Popescu & Lee, 2007; Smith & Lee, 2010), and contributed to changing the misconception that organelle genomes are relatively homogeneous in structure and content (Palmer, 1985; Gray et al., 2004).
Table 1. Examples of the organelle genome architectural diversity among green algae. C = Chlorophyceae; p = prasinophytes; R = charophyte green algae; T = Trebouxiophyceae; U = Ulvophyceae. Genes present in inverted repeats were counted only once.
|
Taxon |
Lineage |
Genome |
% |
% |
# |
# |
Notable |
References |
|
MITOCHONDRIAL |
|
|
|
|
|
|
|
|
|
Ostreococcus tauri |
P |
44.2 |
38.2 |
90 |
65 |
0 |
Compact. |
(Robbens |
|
Pycnococcus provasolii |
P |
24.3 |
37.8 |
87 |
36 |
0 |
Small |
(Turmel |
|
Polytomella capuana |
C |
12.9 |
57.2 |
82 |
10 |
0 |
Linear, |
(Smith |
|
Scenedesmus obliquus |
C |
42.9 |
36.3 |
52 |
42 |
4 |
Deviation |
(Nedelcu |
|
Prototheca wickerhamii |
T |
55.3 |
25.8 |
66 |
61 |
5 |
Non-photosynthetic |
(Wolff |
|
Pedinomonas minor |
T |
25.1 |
22.2 |
58 |
22 |
1 |
Small |
(Turmel |
|
Oltmannsiellopsis viridis |
U |
56.7 |
33.4 |
49 |
54 |
3 |
Repeat |
(Pombert |
|
Pseudendoclonium akinetum |
U |
95.8 |
39.3 |
47 |
57 |
7 |
Expanded |
(Pombert |
|
Chara vulgaris |
R |
67.7 |
40.9 |
52 |
68 |
20 |
Intron |
(Turmel |
|
Chlorokybus atmophyticus |
R |
201.7 |
39.8 |
21 |
70 |
27 |
Among |
(Turmel |
|
PLASTID |
|
|
|
|
|
|
|
|
|
Nephroselmis olivacea |
P |
200.7 |
42.2 |
68 |
128 |
0 |
Large |
(Turmel |
|
Pyramimonas parkeae |
P |
101.6 |
34.7 |
80 |
110 |
1 |
Small |
(Turmel |
|
Floydiella terrestris |
C |
521.1 |
34.5 |
60 |
97 |
26 |
Expanded |
(Brouard |
|
Volvox carteri |
C |
>525 |
~43 |
<20 |
96 |
8 |
Largest |
(Smith |
|
Helicosporidium sp. |
T |
37.4 |
26.9 |
95 |
54 |
1 |
Smallest |
(de |
|
Pedinomonas minor |
T |
98.3 |
34.8 |
71 |
105 |
0 |
Small, |
(Turmel |
|
Bryopsis hypnoides |
U |
153.4 |
33.1 |
37 |
111 |
11 |
Lacks |
(Lü |
|
Pseudendoclonium akinetum |
U |
195.8 |
31.5 |
54 |
105 |
27 |
Large |
(Pombert |
|
Mesostigma viride |
R |
118.3 |
30.2 |
74 |
137 |
0 |
Intron |
(Lemieux |
|
Zygnema circumcarinatum |
R |
165.3 |
31.1 |
55 |
125 |
13 |
Lacks |
(Turmel |
In a broad sense, the organelle genomic architectures of green algae differ from those of land plants (Table 2). Green algae tend to have relatively compact, intron-poor mitochondrial genomes (with some major exceptions), whereas land plant mtDNAs are capacious and intron dense. And green algal plastid genomes are generally larger and more bloated than their mitochondrial counterparts, but for land plants the opposite is true. But as you will see below, it is hard to talk about major trends when dealing with green algal organelle DNAs, as the mitochondrial and plastid genome diversity within green algal groups is often greater than that between groups.
Table 2. Average architectural features of green algal organelle genomes. n, sample size (i.e., number of genomes). For the ranges within groups please refer to GenBank Organelle Genome Resources [www.ncbi.nlm.nih.gov/genome]. For land plant plastid genomes, many of the GenBank entries were incompletely or incorrectly annotated; thus, intron and gene contents were based on the 24 entries for which we were most confident.
|
Phylogenetic Group |
Genome Size (kb) |
% Noncoding |
# of introns |
# of genes |
n |
|
MITOCHONDRIAL |
|
|
|
|
|
|
Chlorophyta |
35.6 |
35.3 |
3.7 |
33 |
19 |
|
prasinophytes |
40.3 |
17.6 |
1 |
57 |
4 |
|
Chlorophyceae |
23.3 |
39.6 |
4.6 |
15 |
10 |
|
Trebouxiophyceae |
43.2 |
37.7 |
3.3 |
47.7 |
3 |
|
Ulvophyceae |
76.3 |
45.9 |
5 |
55.5 |
2 |
|
Charophytes |
92.1 |
47.2 |
15.8 |
67.5 |
4 |
|
Land |
431.2 |
84.3 |
24.4 |
58.0 |
22 |
|
PLASTID |
|
|
|
|
|
|
Chlorophyta |
179.3 |
41.3 |
8.4 |
98.3 |
22 |
|
prasinophytes |
106.9 |
26.2 |
1.5 |
100.2 |
6 |
|
Chlorophyceae |
300.1 |
60.3 |
17.7 |
96.9 |
7 |
|
Trebouxiophyceae |
116.9 |
31.6 |
1.8 |
96.0 |
6 |
|
Ulvophyceae |
167.1 |
49.4 |
16.0 |
104.5 |
3 |
|
Charophytes |
151.5 |
41.2 |
9.7 |
128.8 |
6 |
|
Land |
147.8 |
41.9 |
23.8 |
106.9 |
107a |
Although organelle genomes tend to be rich in adenine and thymine (AT-rich), data from certain green algae have shown that this is not always the case. The mtDNA of the non-photosynthetic, unicellular chlamydomonadalean Polytomella capuana is 57% guanine and cytosine (GC), making it one of only a few GC-biased organelle genomes sequenced to date (Smith & Lee, 2008). Other green algae also harbour GC-rich organelle DNA: Analyses of the mtDNA-encoded cox1 and/or cob from Oogamochlamys gigantea, Lobochlamys segnis, and Lobochlamys culleus (all from the Oogamochlamydinia-clade, sensu Nakada et al. (2008a)) revealed average GC compositions of 50, 54, and 63%, respectively, suggesting that the overall mitochondrial GC content of at least some of these species is higher than that of P. capuana (Borza et al., 2009); and the unpublished mitochondrial and plastid genome sequences of the trebouxiophyte Coccomyxa sp. C-169 (GenBank accession numbers HQ874522 and HQ693844) are 53% and 51% GC, respectively — only the second example (the other being the lycophyte Selaginella moellendorffii (Smith, 2009)) of a species with GC-biased DNA in both the mitochondrial and plastid compartments. In land plants, the levels of G and C in the mitochondrial and plastid genomes are positively correlated with the number of C-to-U RNA editing sites (Malek et al., 1996; Jobson & Qiu, 2008; Smith, 2009), but this is not the case for green algae as there is no evidence of RNA editing in either their mitochondrial or plastid genomes (Lenz et al., 2010, and references therein). Most green algal organelle DNAs have a more “typical” nucleotide composition, showing a propensity for A and T, sometimes strongly so, as in the cases of the Chara vulgaris, Scenedesmus obliquus, Helicosporidium sp., and Leptosira terrestris plastid genomes, which are among the most AT-biased ptDNAs sequenced to date (de Cambiaire et al., 2006; de Koning & Keeling, 2006; Turmel et al., 2006a). Whether the nucleotide bias is towards A and T or G and C, it is usually expressed most strongly within green algal organelle DNAs at what are considered to be among the more neutrally evolving sites in a genome (i.e., intergenic and intronic positions and the synonymous sites of protein-coding DNA), implying that the forces driving green algal organelle nucleotide landscape are nonadaptive (Smith & Lee, 2008; Borza et al., 2009).
It was once assumed that all mitochondrial genomes are circular (or at least circular-mapping) molecules. However, it is now well established that linear and linear fragmented mtDNAs have evolved numerous times in diverse eukaryotic lineages (Nosek & Tomáska, 2003), including the Reinhardtinia clade (sensu Nakada et al., 2008a) of the Chlamydomonadales. Every Reinhardtinia-clade mtDNA examined thus far appears to have either a linear monomeric or a linear fragmented conformation (Laflamme & Lee, 2003; Mallet & Lee, 2006), with the exception of the Volvox carteri mtDNA, which assembles as a circular molecule (Smith & Lee, 2010). Conversely, all of the characterized green algal mitochondrial genomes from outside this clade map in genome assemblies and/or gel electrophoresis studies as unit-sized circular chromosomes save for the mtDNAs of some Lobochlamys taxa, which may be linear fragmented (Borza et al., 2009). Studies on linear green algal mtDNAs have revealed a range of interesting telomeric sequences and structures, including inverted repeats, 3´ overhangs, and closed single-stranded loops (Vahrenholz et al., 1993; Smith & Lee, 2008). Since mitochondria lack telomerase, it is presumed that the elaborate termini of linear mtDNAs help the genome overcome the end replication problem, as defined by Olovnikov (1971) and Watson (1972). All of the available ptDNA sequences for green algae assemble as genome-sized circular molecules, but, like the mitochondrial and plastid DNAs from land plants, they probably exist in vivo as multi-genome-sized branched linear forms, which can (as a byproduct of recombination-dependent replication) recombine to generate unit-sized circular molecules (Simpson & Stern, 2002; Bendich, 2004; 2007). Recent data from the ptDNA of the ulvophyte Bryopsis hypnoides support this hypothesis (Lü et al., 2011).
Green algal organelle DNAs vary in size and compactness (Tables 1 and 2). The smallest and most reduced plastid genome observed from a green alga belongs to the non-photosynthetic parasite Helicosporidium sp. (Trebouxiophyceae) — it is 37.4 kb and 95% coding DNA (de Koning & Keeling, 2006). In contrast, the plastid genome of the free-living photosynthetic trebouxiophyte Leptosira terrestris is 195 kb, half of which is noncoding DNA (de Cambiaire et al., 2007). Compact plastid genomes are also found in some prasinophytes, including Ostreococcus tauri, Micromonas spp., and Pycnococcus provasolii whose ptDNAs are around 75 kb and 80% coding (Robbens et al., 2007a; Turmel et al., 2009a). But this is not a trend of the entire genus: the Nephroselmis olivacea ptDNA is 201 kb, a third of which represents noncoding nucleotides (Turmel et al., 1999b). The plastid genomes of the chlorophyceans Floydiella terrestris and Volvox carteri are ~525 kb and about 80% noncoding DNA (Brouard et al., 2010; Smith & Lee, 2010), making them almost 300 kb larger than any other available ptDNA sequence. Large plastid genomes are a common theme among chlorophycean algae: the sequenced ptDNAs from C. reinhardtii, Stigeoclonium helveticum, and Dunaliella salina are 204, 223, and 269 kb, respectively (Maul et al., 2002; Bélanger et al., 2006; Smith et al., 2010b), and gel electrophoresis results place the plastid genomes of both Chlamydomonas gelatinosa and Chlamydomonas moewusii at ~290 kb (Boudreau et al., 1994; Boudreau & Turmel, 1996). But the largest green algal plastid genomes probably come from certain Acetabularia species (Ulvophyceae), which are believed to have ptDNAs in excess of 2 Mb (reviewed by Palmer, 1985).
The mtDNAs from green algae can also be large, although they are generally smaller than their plastid counterparts and much smaller than land plant mtDNAs, which can achieve sizes of 3 Mb (Ward et al., 1981). The charophyte Chlorokybus atmophyticus has the largest recorded green algal mitochondrial genome (201 kb and 77% noncoding) — more than twice the size of any other sequenced green algal mtDNA (Turmel et al., 2007a). It closest rivals in this respect are the mitochondrial genomes of Chara vulgaris (67.7 kb, 47.5% noncoding) and the ulvophyte Pseudendoclonium akinetum (96 kb, 50% noncoding) (Turmel et al., 2003; Pombert et al., 2004). The most diminutive mtDNAs from the Viridiplantae are currently found in the Chlorophyceae. For example, the mtDNAs of Polytomella species range from 13-16 kb (Smith et al., 2010a) and that of C. reinhardtii varies from 16-19 kb (depending on the presence of optional introns) (Gray & Boer, 1988; Michaelis et al., 1990; Vahrenholz et al., 1993). Though small, chlorophycean mtDNAs can be quite bloated; both the D. salina and V. carteri mtDNAs are ~60% noncoding (Smith et al., 2010b). Trebouxiophytes and charophytes can also have small mtDNAs: those of Pedinomonas minor and M. viride are 25 and 42 kb, respectively (Turmel et al., 1999a; Turmel et al., 2002c). Prasinophytes tend to have the most close-packed mitochondrial genomes within the Viridiplantae, as exemplified by O. tauri, Micromonas spp., and P. provasolii, whose mtDNAs are between 82-92% coding (Robbens et al., 2007a; Turmel et al., 2008); this compact architecture is paralleled in their nuclear and plastid compartments (Derelle et al., 2006; Worden et al., 2009).
As alluded to above, differences in the amount of intronic and intergenic DNA are responsible for much of the variability in mitochondrial and plastid genome size observed among green algae. The number of introns (group I and group II) in green algal organelle genomes varies for mtDNA from 0 (Polytomella and most prasinophyte species) to 27 (C. vulgaris) and for ptDNA from 0 (M. viride, Micromonas spp., N. olivacea, and P. minor) to maximums of 26, 27, and 36 (F. terrestris, P. akinetum, and D. salina, respectively). Intergenic sequences can account for as little as 5% (e.g., the ptDNA of Helicosporidium sp.) to more than 50% (e.g., the ptDNAs of V. carteri, L. terrestris, and C. reinhardtii, and the mtDNA of C. atmophyticus) of green algal organelle genomes. In many cases the intergenic regions (and sometimes the intronic DNA) of green algal mitochondrial and plastid genomes are overrun with repetitive elements (Maul et al., 2002; Pombert et al., 2005; Smith & Lee, 2009). Green algal organelle DNA repeats come in a range of sizes, orientations (e.g., tandem, palindromic, inverted), nucleotide compositions, and complexities, and can often be folded into secondary structures such as hairpin loops. Some are believed to be transposable elements, and there are examples of similar types of repeats being found in both the mitochondrial and plastid compartments of the same species (Pombert et al., 2005; Smith & Lee, 2009). It has also been argued that in some green algal lineages organelle repeats have played a role in fashioning the mitochondrial and plastid genomes by acting as catalysts for genome fragmentation and reorganization as well as gene splitting and scrambling (Boer & Gray, 1991; Nedelcu, 1998; Nedelcu & Lee, 1998; Maul et al., 2002). Moreover, certain repeats, especially those that can be folded into secondary structures, are thought to be involved in gene regulation and processing (Boer & Gray, 1986; Jiao et al., 2004; Smith & Lee, 2008).
Most plastid genomes have a “quadripartite structure” where a single set of large inverted repeats, which typically contain the rRNA-coding genes and various other genes, divide the genome into two single-copy regions. The quadripartite structure is believed to have been present in the plastid genome of the ancestor that gave rise to green plants, and departure from it is relatively rare among plastid-harbouring eukaryotes. Thus, it is significant that there are examples from each of the major green algal groups of species that have lost (or almost lost) their ptDNA inverted repeats. Examples include the charophyte green algae Staurastrum punctulatum and Zygnema circumcarinatum (Turmel et al., 2005), the chlorophyceans F. terrestris and S. helveticum (Bélanger et al., 2006; Brouard et al., 2010), the prasinophytes P. provasolii and Monomastix (Turmel et al., 2009a), the ulvophyte B. hypnoides (Lü et al., 2011), and the trebouxiophytes L. terrestris and Helicosporidium sp. (de Koning & Keeling, 2006; de Cambiaire et al., 2007). These observations suggest that loss of the inverted repeat has occurred multiple times throughout the evolution of green algae — and sometimes multiple times within the same group, as in the case of trebouxiophytes (de Cambiaire et al., 2007). Interestingly, the O. tauri mtDNA has an inverted repeat region within its mitochondrial genome, which, like that of plastid genomes, harbours the rRNA-coding genes as well as other types of genes (Robbens et al., 2007a).
Some green algal organelle genomes abound with genes whereas others are gene depauperate. The available green algal ptDNAs contain anywhere from around 55-135 genes, typically representing 3 rRNAs (rrnS, rrnL, and rrnF), 25-35 tRNAs, and approximately 25-100 proteins, which are involved in processes such as photosynthesis, metabolism, transcription, and translation. The sequenced mtDNAs from green algae contain around 10-70 genes, which encode 2-3 rRNAs (rrnS, rrnL, and, sometimes, rrnF), approximately 1-28 tRNAs, and around 7-37 proteins, most of which function in transcription, translation, or oxidative phosphorylation. Charophyte green algae tend to have gene-abundant mitochondrial and plastid genomes, as demonstrated by C. atmophyticus and C. vulgaris, which have 70 and 68 mitochondrial-encoded and 138 and 127 plastid-encoded genes, respectively (Turmel et al., 2006a; Lemieux et al., 2007). The organelle genomes from chlorophyceans, particularly the mtDNAs of chlamydomonadalean algae, are gene poor. For instance, the C. reinhardtii mitochondrial and plastid genomes contain 13 and 94 genes, respectively (Gray & Boer, 1988; Maul et al., 2002), and the mtDNAs from Polytomella species (which are the most reduced mtDNAs observed from green plants) encode only seven proteins, two rRNAs, and one tRNA (Fan & Lee, 2002; Smith et al., 2010a). Other examples of reduced organelle gene contents include the Helicosporidium sp. ptDNA, which contains no genes for photosynthetic proteins, and the P. minor mtDNA, which has 22 genes, none of which code for ribosomal proteins.
Studies on the organization and architecture of genes within green algal organelle genomes — including their arrangement and conservation among species, intron content and insertion sites, genomic localities (i.e., inside or outside of repeated regions), and sequence divergence — have helped infer the phylogenetic relationships among green plants and provided insights into the modes and tempos of green algal genome evolution. Some interesting findings to have come from these investigations include observed changes from the standard genetic code. In the S. obliquus mitochondrial genome TCA (normally a serine codon) is a stop codon and TAG (normally a stop codon) codes for leucine (Nedelcu et al., 2000), and in the P. provasolii mtDNA TGA (normally a stop codon) codes for tryptophan and TTA and TTG (normally leucine codons) are stop codons (Turmel et al., 2010). Other discoveries include the presence of fragmented and scrambled small- and/or large-subunit rRNA-coding genes in the mtDNAs from chlorophycean species and P. minor (Gray & Schnare, 1996; Nedelcu, 1997; Turmel et al., 1999a). Fragmented protein-coding genes, which are brought together at the RNA level via intron trans-splicing, have been identified in both the mitochondrial and plastid genomes of green algae (Glanz & Kuck, 2009; Pombert & Keeling, 2010). Genes acquired through horizontal gene transfer have been reported. The int and dpoB genes, which are located in the ptDNA inverted repeats of O. cardiacum, and putatively code for a tyrosine recombinase and a DNA-directed DNA polymerase, are believed to have been obtained laterally, possibly from the mitochondrial plasmid of a fungus (Brouard et al., 2008). Also, the Oltmannsiellopsis viridis mtDNA has what appears to be a recently captured integrase gene originating from a bacterium and a group II intron coming from a cryptophyte (Pombert et al., 2006), the P. parkeae ptDNA harbors a DNA primase gene putatively acquired from a virus (Turmel et al., 2009a), and the mitochondrial genome of C. reinhardtii contains a reverse-transcriptase-like gene (rtl) whose evolutionary origins and function remain unknown (Boer & Gray, 1988).
Much effort has been spent in trying to understand the evolutionary forces responsible for the architectural variation among green algal organelle DNAs. One hypothesis that has been tested on multiple fronts is the mutational-hazard hypothesis, which suggests that the primary forces governing organelle genome size and structure are mutation and random genetic drift (Lynch & Conery, 2003; Lynch et al., 2006). The hypothesis argues that genomic embellishments, such as introns and intergenic DNA, are a mutational liability because they represent targets for potentially deleterious mutations, where the higher the mutation rate the greater the burden of the embellishment. It is further argued that species with large effective genetic population sizes (i.e., large Ne) are more efficient at perceiving and eliminating burdensome DNA than those with small effective population sizes. Two types of silent-site DNA sequence divergence data — that within species (πsilent) and that between species (dS) — have provided clues into the roles that Ne and the ratio of the per-generation rate of mutation per nucleotide site (µ) have had in shaping green algal organelle genome structure. Available πsilent measurements (which can be used as a proxy for Neμ) from V. carteri are on average very low (around 0.0004, 0.0006, and 0.005 for the mtDNA, ptDNA, and nucDNA, respectively), especially relative to other protists (Smith & Lee, 2009; Smith & Lee, 2010). This suggests that V. carteri has a small Neμ and therefore a reduced ability to detect and eradicate excess DNA, which could help explain why its organelle and nuclear genomes are among the most inflated from green algae. Studies on the relative dS values (which can be used to estimate relative mutation rates) from Chlamydomonas and Mesostigma indicate that in each of these algae the mutation rates of the mitochondrial, plastid, and nuclear compartments are similar (Popescu & Lee, 2007; Hua et al., 2011). This is in sharp contrast to the situation for most land plants where the silent-site substitution rate of the mtDNA is estimated to be three-fold lower than that in the ptDNA and 12-times lower than that in the nuclear DNA (Wolfe et al., 1987; Wolfe et al., 1989). This may help explain why, broadly speaking, there is less disparity in genomic complexity between the mitochondrial and plastid genomes of green algae than there is for land plant mitochondrial and plastid DNAs. The average fraction of noncoding DNA in the available mtDNA and ptDNA sequences from chlorophytes is 0.35 and 0.41, respectively, and for charophyte green algae the corresponding values are 0.47 (mtDNA) and 0.41 (ptDNA). Conversely, the proportion of noncoding nucleotides in the mtDNA of land plants averages 0.84 but is 0.42 for the ptDNA (Table 2). Thus, the comparable architectures of green algal plastid and mitochondrial DNAs may be a consequence of these genomes having similar mutation rates, as reflected in their similar rates of silent-site substitution. As more green algae organelle DNA data become available — especially data from different members of the same population and from closely related species — we will get a better idea of now Ne and μ are shaping green algal genome architecture.
B. Ecology and molecular evolution of oceanic picoplanktonic prasinophytes
The marine planktonic compartment of picoeukaryotes has been previously overlooked because of the extremely small size of its constituents (1-3µm). However, its photosynthetic representatives, although present in lower concentration than cyanobacteria, are metabolically very active and can represent up to 80 % of the biomass primary production in coastal ecosystems (Li, 1994; Worden et al., 2004). Picoeukaryote composition is highly diverse, encompassing several main branches of the eukaryotic crown (Lopez-Garcia et al., 2001; Moon-van der Staay et al., 2001; Vaulot et al., 2008), and, among this diversity, the prasinophytes are constantly present and emerged as a major component, especially in coastal areas (Diez et al., 2001; Massana et al., 2004; Not et al., 2004; Piganeau & Moreau, 2007; Viprey et al., 2008). The prasinophytes form a paraphyletic assemblage of at least nine clades, with the Mamiellophyceae (prasinophyte clade II) being common in mesotrophic coastal areas (Zhu et al., 2005; Countway & Caron, 2006; Marie et al., 2006) and clades VIII and IX (known exclusively from environmental sequencing) in oligotrophic oceanic waters (Lepère et al., 2009; Shi et al., 2009). Sequences belonging to the Picocystis clade (prasinophyte clade VII) correspond to strains originally isolated from open oceanic areas but are also encountered in clone libraries originating from coastal regions (Shi et al., 2009).
Micromonas pusilla (Butcher, 1952) was the first species to be discovered among the Mamiellophyceae. Since this date, at least seven new genera of this class have been discovered over the last decade by electron microscopic and phylogenetic analyses (Marin & Melkonian, 2010), currently giving a total of 19 species. Most of the clone library studies showed that among Mamiellophyceae, the genera Bathycoccus, Micromonas and Ostreococcus are dominant in many different areas (Guillou et al., 2004; Not et al., 2004; Marie et al., 2006; Viprey et al., 2008). These three genera grow well in culture and are also well-represented in culture collections. Phylogenies based on the 18S rDNA showed that Micromonas and Ostreococcus are divided in several clades (Guillou et al., 2004) corresponding to different ecotypes and probably different species (see below) (Rodriguez et al., 2005; Šlapeta et al., 2006). This has been shown on the basis of physiological properties (Rodriguez et al., 2005; Cardol et al., 2008), geographical origin (Foulon et al., 2008), and niche partitioning (Demir-Hilton et al., 2011) of the strains/ecotypes. For example, “low light” and “high light” ecotypes have been described in Ostreococcus, corresponding to different clades (Rodriguez et al., 2005). A recent study showed that co-occurrence of both ecotypes at the same geographical location is rare, although parameters explaining clade distribution are more complex than irradiance alone (Demir-Hilton et al., 2011). It is proposed that these two “low light” and “high light” ecotypes could rather correspond to oceanic and coastal clades/ecotypes, respectively. In contrast, all Bathycoccus sequences are grouped in a homogeneous clade (Guillou et al., 2004), even with the addition of new strains isolated recently from different areas (Moreau, unpublished). The ecological importance of Bathycoccus is probably overlooked (Johnson & Sieburth, 1982; Eikrem & Throndsen, 1990), but it is sporadically found in clone library studies where it can even be dominant (Marie et al., 2006).
The wide distribution of Mamiellophyceae and especially the three genera mentioned above, their ecological importance, ease of culturing and small genomes have made these eukaryotic micro-organisms emerging models for ecological and biological studies of marine phytoplankton. Two Micromonas (Worden et al., 2009) and two Ostreococcus (Derelle et al., 2006; Palenik et al., 2007) genome sequences have been published, and the genome sequence of a low light strain of Ostreococcus is also available on the JGI web site (genome.jgi-psf.org). These genome sequences allowed a quantification of the genetic distances between the different clades inside a genus. For example, in two sequenced Ostreococcus strains belonging to two different clades, the 18S rDNA divergences was less than 0.5% whereas around 25% divergence in amino-acid identity over their orthologous protein coding genes was observed. An even higher divergence was observed between the two sequenced Micromonas strains (Worden et al., 2009). This high genome sequence divergence and a different number of chromosomes, are not compatible with genetic exchange via meiosis, and have led to the “genomic species” definition of O. lucimarinus (Palenik et al., 2007). Sexual reproduction has never been observed between any Mamiellophyceae strains, although a complete gene set necessary to control meiosis is present in both Micromonas and Ostreococcus (Derelle et al., 2006; Worden et al., 2009). However, evidences of recombination events in several Ostreococcus tauri strains strongly suggest genetic exchanges among strains belonging to the same clade (Grimsley et al., 2010).
The five available Mamiellophyceae genomes are very gene-dense, with protein-coding sequences representing almost 70% of the genome for most strains. Their small cellular and genome sizes have led to the hypothesis that they may represent the “bare limits” of life as a free-living photosynthetic eukaryote, presumably having disposed of redundancies and presenting a simple organization. Indeed, for most of the annotated metabolic processes, a reduction in the number of gene copies was observed, usually down to a single copy. Complete genome sequences have provided insights into potential Mamiellophyceae metabolisms and adaptations, although most of them still await experimental confirmation. For example, biochemical studies have shown that the same light-harvesting chlorophyll-binding proteins (LHCs) were associated with both PSI and PSII. However, the availability of Micromonas and Ostreococcus genomes confirmed the existence of an unusual Mamiellophyceae-specific LHC, probably associated with both photosystems (Cardol et al., 2008; Six et al., 2009), but also revealed the presence of specific PSI and PSII subunits, which are expressed at low levels and which were not detected biochemically (Six et al., 2005). Because Mamiellophyceae are considered as early-diverging green algae, these findings had important evolutionary implications and supported the hypothesis of an ancient origin of LHCI genes from which LHCII and the major Mamiellophyceae LHC genes probably evolved, consolidating data into a coherent evolutionary scenario. Likewise, genes encoding all of the enzymes required for C4-photosynthesis are present in both the Micromonas and Ostreococcus genomes indicating that, like diatoms, these algae may use this pathway as a CO2 concentration mechanism (Derelle et al., 2006; Worden et al., 2009). This may, therefore, be a strategy general to planktonic microalgae, raising the possibility that, despite its high energetic cost, C4-photosynthesis might give a critical ecological advantage in CO2-limiting conditions, such as in phytoplankton blooms.
Other examples of potential metabolic adaptations can be found in the available genomes. For example, in unicellular eukaryotes a general system for iron (Fe) uptake usually involves a ferric reductase, multicopper oxidase, and a ferric permease. Ostreococcus, in contrast, appears to lack all of these iron transport components, with the possible exception of a multicopper oxidase found only in O. tauri (Palenik et al., 2007). This implies that this genus has a different system of Fe acquisition compared to those of major competitors. Furthermore, Ostreococcus shows several adaptations that reduce Fe requirements, such as the absence of Cytochrome c6 in Ostreococcus and its replacement by plastocyanin as the only electron carrier between the Cyt b6/f complex and photosystem I (Palenik et al., 2007). In silico analysis of nutrient acquisition has also been investigated by comparative analysis of genes involved in nitrogen metabolisms. Eight of the genes involved in nitrate uptake and assimilation, as well as four genes for urea assimilation are found next to each other in two clusters located on two chromosomes in Ostreococcus (Derelle et al., 2006; Palenik et al., 2007). A comparable clustering of nitrate assimilation genes is also observed in Micromonas, whereas in C. reinhardtii fewer of these genes are clustered. This organization indicates a possible selective pressure for optimization of nitrate and urea uptake and assimilation in microalgae. A recent study in Micromonas showed a high degree of “mixed lineage gene affiliations”, for nitrogen transport and assimilation genes (McDonald et al., 2010), suggesting ancient origin, complex evolution pattern of gene duplication and gene loss, or horizontal gene transfer for some of these genes.
These in silico analyses are still in their infancy and many metabolisms, potential adaptations and/or evolutive signatures still remain to be elucidated. Beside portals dedicated to the annotation of each of the different genomes, web sites focused on comparative analysis between the different organisms belonging to the green lineage, as PLAZA (bioinformatics.psb.ugent.be/plaza/) (Proost et al., 2009) and its derivative Pico-PLAZA are now available. Such tools for comparative approaches, coupled to the development of new sequencing possibilities for both new genomes and metagenomes (Cheung et al., 2010; Marie et al., 2010) from various environments open the door to a better understanding of the adaptations of the organisms to their environment.
C. Genomic insights into the evolution of complexity in volvocine green algae
The volvocine algae (Chlorophyceae: Chlamydomonadales) are an important model system for understanding the evolution of multicellularity and cellular differentiation. This informal grouping, imprecisely defined as “Volvox and its close relatives”, is usually taken to include the families Volvocaceae, Goniaceae and Tetrabaenaceae (Nozaki et al., 2000; Nozaki et al., 2003) along with a few unicells in the genera Chlamydomonas and Vitreochlamys. Their appeal as a model system stems from the relative simplicity of their development and from the wide range of sizes and degrees of complexity found in extant species. Cell numbers range from 1 to ~50,000, including nearly every power of 2 (4, 8, 16, etc.) in this range, and multicellular species may have partial, complete, or no differentiation between reproductive and somatic cells (Fig. 5).

Two species in particular, the unicellular Chlamydomonas reinhardtii and the differentiated multicellular Volvox carteri forma nagariensis (V. carteri hereafter), have been the focus of molecular-genetic investigations, and their genomes have recently been sequenced (Merchant et al., 2007; Prochnik et al., 2010). C. reinhardtii is a biflagellate, facultatively heterotrophic soil alga that serves as an important model organism for understanding eukaryotic photosynthesis, flagellar motility, and basal body function. V. carteri is a spheroid typically made up of ~1000-2000 small, biflagellate somatic cells, ~8-16 much larger reproductive (germ) cells, and a gelatinous extracellular matrix (ECM) that makes up ~99% of the total volume. Several of the genes underlying multicellular development and cellular differentiation in V. carteri have been identified and cloned (Kirk, 1998; Kirk, 2005), greatly improving our understanding of the molecular and developmental bases of these traits.
In spite of the dramatic differences in size and complexity between C. reinhardtii and V. carteri, their genomes are similar in size and gene content. The V. carteri nuclear genome is ~17% larger than that of C. reinhardtii (138 Mb vs. 118 Mb), but this difference is largely due to non-coding sequences, as V. carteri has greater repeat content and, on average, longer introns (Prochnik et al., 2010). The total number of genes is similar, with ~14,500 genes in each species’ genome (Merchant et al., 2007; Prochnik et al., 2010). Most gene families have similar numbers of genes in each species, although several interesting exceptions occur (discussed below).
Since its divergence from unicellular ancestors ~230 million years ago (Herron et al., 2009), the lineage leading to V. carteri has undergone a series of developmental changes resulting in a multicellular organism with a distinct anterior-posterior polarity, a complex developmental program, and a clear distinction between reproductive and somatic cells. The evolutionary path of V. carteri from a single-celled ancestor was mapped by Kirk (2005), who divided the evolution of multicellularity and cellular differentiation into a series of twelve developmental changes. For several of these changes, information about the underlying genetics is available, and there are several genomic differences that suggest possible genes whose roles have yet to be identified.
In C. reinhardtii, and presumably in the unicellular ancestor of V. carteri, cell division occurs through palintomy rather than by binary fission; that is, cells grow to many times their original size without dividing and then undergo several rounds of rapid cell division with little or no growth between successive divisions (Sleigh, 1989). In C. reinhardtii, the resulting offspring emerge from the mother cell wall and begin their lives as separate unicellular organisms. Most multicellular volvocine algae have retained this mode of cell division, producing large reproductive cells, each of which divides n times to produce 2n daughter cells (n ranges from 2 to ~14). In the multicellular volvocine algae, the daughter cells resulting from palintomic division of a reproductive cell remain connected to form a single, multicellular offspring.
The first step in the evolution of multicellularity in the volvocine algae was probably the origin of some means of keeping daughter cells connected to each other. The mechanisms by which this attachment occurs differ among volvocine species. In the Tetrabaenaceae, the smallest multicellular volvocine algae and the earliest to diverge from the lineage leading to V. carteri, palintomic division produces four daughter cells that remain embedded in a common ECM (Iyengar & Desikachary, 1981). In all other species, cells are held together by specialized cell wall attachments or by a boundary layer that surrounds the entire organism (Nozaki, 1990; Nozaki & Kuroiwa, 1992). Both the ECM and the boundary layer are homologs of portions of the C. reinhardtii cell wall (Kirk et al., 1986).
The ECM in V. carteri is a complex, multifunctional structure that makes up over 99% of the volume of mature spheroids (Hallmann, 2003). Two gene families involved in ECM synthesis and function are substantially larger in the V. carteri genome than in that of C. reinhardtii: the pherophorin family has 49 members in V. carteri and only 22 in C. reinhardtii, while the Volvox metalloproteinase family has 42 members in V. carteri and only 8 in C. reinhardtii (Prochnik et al., 2010). Expansion of these gene families may thus have been an important factor in the increase in size and complexity of ECM in multicellular volvocine algae. The volume of ECM has undergone several expansions and contractions throughout volvocine evolution (Herron & Michod, 2008), and it would be interesting to know if the same genes were involved in lineages other than that leading to V. carteri.
Several of the important differences between single-celled and multicellular volvocine algae can be attributed to differences in cell cycle regulation. Thanks to the palintomic cell division program, and specifically to the lack of growth between cell divisions, the reproductive cells of large multicellular species must postpone the start of cell division until they are large enough to produce all of the cells in the offspring. In addition, early in development the cells of multicellular species are connected by cytoplasmic bridges resulting from incomplete cytokinesis (Kirk, 2005). In most species these connections break down before maturity (Stein, 1965; Marchant, 1977; Fulton, 1978). Most of the genes involved in cell cycle regulation have one ortholog each in V. carteri and C. reinhardtii, but one family of cell cycle-related proteins, the D cyclins, is larger in V. carteri (Prochnik et al., 2010). Prochnick et al. (2010) suggest that the additional members of this gene family may play a role in the differences in timing and extent of cell division between single-celled and multicellular volvocine algae.
One of the important functions of the cytoplasmic bridges that result from incomplete cytokinesis is in the process of inversion, a developmental process that occurs in all members of the Volvocaceae (Green et al., 1981). In this family, at the end of cell division, embryos are cup- or bowl-shaped, but with their flagella on the inner (concave) surface (Gerisch, 1959; Marchant, 1977; Fulton, 1978; Kirk, 2005). During inversion, cells move relative to the cytoplasmic bridges in order to turn the embryo inside-out, so that the flagella end up on the outer surface (Gerisch, 1959; Marchant, 1977; Fulton, 1978; Green et al., 1981). This process seems to have evolved by co-option of genes already present in the unicellular ancestor of V. carteri: three genes known to be involved in V. carteri inversion (invA, invB, invC) have highly conserved orthologs in C. reinhardtii (Nishii et al., 2003; Ueki & Nishii, 2009; Nishii & Miller, 2010), and an invA ortholog (IAR1) in C. reinhardtii is capable of rescuing V. carteri invA mutants (Nishii et al., 2003).
Changes in cell cycle regulation are also involved in cellular differentiation. The distinction between somatic and reproductive cells in V. carteri is established early in development, when different numbers and patterns of divisions – including asymmetric divisions – produces cells of two different sizes (Starr, 1969). The ~30-fold difference in cell size triggers two distinct patterns of gene expression that establish the fate of the large and small cells as gonidia (asexual reproductive cells) and somatic cells, respectively (Kirk & Kirk, 1985; Tam & Kirk, 1991; Kirk et al., 1993). The genes involved in asymmetric division and the differentiation of somatic cells have been subjected to intensive study, and the overall picture seems to be one of co-option of genes already present in the unicellular ancestor. For example, the glsA (gonidialessA) gene, which is required for asymmetric division in V. carteri, has a homolog in C. reinhardtii that is capable of rescuing V. carteri glsA mutants (Cheng et al., 2003). The regA (somatic regenerator) gene, a member of the VARL (Volvox algal regA-like) family, suppresses growth and reproduction in V. carteri somatic cells (Kirk et al., 1999; Meissner et al., 1999; Duncan et al., 2007). Although C. reinhardtii lacks an exact regA ortholog, phylogenetic analysis of the VARL gene family suggests that this is due to gene loss in the C. reinhardtii lineage rather than a gene duplication in the V. carteri lineage (Duncan et al., 2007). The most closely related C. reinhardtii regA homolog is induced under nutrient and light-deprivation, leading to the suggestion that the somatic differentiation function of regA was co-opted from an environmental acclimation function in the unicellular ancestor of V. carteri (Nedelcu & Michod, 2006; Nedelcu, 2009).
Another form of cellular differentiation is between the dissimilar gametes of anisogamous (or oogamous) mating types. Both C. reinhardtii and V. carteri are facultatively sexual and heterothallic (sex or mating type is genetically determined), but their sexual phase differs in several respects. In C. reinhardtii, entry into the sexual phase of the life cycle is induced by nitrogen starvation, which causes haploid vegetative cells to differentiate into isogamous gametes (Sager & Granick, 1954). Two gametes of opposite mating type fuse to form a diploid zygote, which undergoes meiosis to produce four haploid vegetative cells. In V. carteri, sex is induced by a sex-inducing pheromone that can be either produced by the somatic cells of haploid asexual spheroids in response to heat stress or released from the sperm packets of spontaneously occurring sexual males (Kirk & Kirk, 1986). Asexual spheroids respond to the sexual inducer by producing male or female offspring, which in turn produce either eggs or packets of 64 or 128 sperm (Kirk, 1998). Sperm and eggs fuse to form a dormant diploid zygote, which undergoes meiosis to form a single asexual spheroid and three polar bodies (Kirk, 1998).
In C. reinhardtii, mating type is determined within the ~200 kb MT region, in which meiotic recombination is suppressed by differences in gene order (Ferris & Goodenough, 1994; Ferris et al., 2002). Within this region, the MID (Minus Dominant) gene, found only in the minus mating type, acts as a dominant Mendelian trait determining mating type (Ferris & Goodenough, 1997). Discovery of a MID ortholog in males of Pleodorina starrii, an oogamous volvocine alga, showed that males evolved from the minus mating type; this is the first case in which homology between an oogamous sex and an isogamous mating type has been demonstrated (Nozaki et al., 2006a). As in the C. reinhardtii MT region, the V. carteri MT region includes genes that are shared between both sexes as well as several that are limited to either males or females (Ferris et al., 2010). However, the V. carteri MT region is around five times as large, includes substantially more genes, and has, on average, much greater sequence divergence between alleles of genes found in both sexes (Ferris et al., 2010). Most of the sex-limited genes are differentially expressed during sexual differentiation, and many of the shared genes are differentially expressed both during sexual differentiation and between males and females (Ferris et al., 2010). Differential expression, along with the increased gene content of the V. carteri MT region relative to that of C. reinhardtii, suggests a number of candidate genes that could have been important for the evolution of oogamy.
Although V. carteri, with just two cell types, is often cited as an example of a simple multicellular organism, its structure and development have changed a great deal relative to its unicellular ancestors. Aside from a 2000-fold increase in cell number and a million-fold increase in total volume, the lineage leading to V. carteri has undergone several important changes in cell cycle regulation resulting in the establishment of cytoplasmic connections among cells and of a reproductive-somatic division of labor. Changes in flagellar arrangement and a coordinated developmental process of inversion produced a multicellular organism with a distinct anterior-posterior polarity and the ability to coordinate the movements of 4000 flagella in order to seek out optimal light levels (Hoops, 1993). Changes in the sexual phase of the life cycle have led to the establishment of males and females from isogamous ancestors, and entry into the sexual phase has become dependent on signals from other individuals rather than on direct sensing of environmental conditions.
Comparison of the V. carteri and C. reinhardtii genomes has already revealed much, including some surprises, about the evolution of complexity in the volvocine algae. In a few cases, differences in the sizes of gene families suggest that gene duplication has played a role in the evolution of multicellular development and cellular and sexual differentiation. In most cases, though, the traits that comprise a differentiated multicellular organism seem to be controlled by genes co-opted from a unicellular ancestor. Even in those cases in which genomic comparisons suggest candidate genes, their specific functions have yet to be identified, nor, for most, has their involvement in particular traits even been confirmed. Furthermore, with few exceptions, we know nothing about the genetic basis of most traits in any of the ~50 volvocine species other than V. carteri and C. reinhardtii. Publication of the V. carteri genome has opened up the comparative genomics phase of volvocine research, suggesting answers to some questions and posing new ones that could not previously have been framed. Thus volvocine genomics is likely to accelerate the pace of discovery and to continue producing fruitful research for some time to come.
D. Genetic codes and the translational apparatus in green seaweeds
Translation is the first step of protein biosynthesis in which the ribosome decodes messenger RNA (mRNA) to form polypeptides that will later fold into the protein’s active structure. The eukaryotic ribosome consists of two subunits, each of which is composed of a folded RNA and several ribosomal proteins. A crucial aspect in translation is the recognition of the codons in the mRNA by anticodons on the transfer RNAs (tRNA), which carry the corresponding amino acids. This recognition is generally well-conserved across eukaryotes, leading to a nearly universal genetic code linking codons to their corresponding amino acids.
Deviations from the canonical genetic code are known in some eukaryotes, and they have also been shown in some green algal groups. Early work on cDNA of the nuclear-encoded small subunit of the Rubisco gene showed that the stop codons UAA and UAG were reassigned to code for glutamine in the siphonous dasycladalean green seaweeds Acetabularia and Batophora (Schneider et al., 1989; Schneider & Degroot, 1991). More recently, the evolution of these codon reassignments among the Ulvophyceae was studied in more detail using multi-gene datasets (Gile et al., 2009; Cocquyt et al., 2010a). These studies have shown a complex distribution of the alternative genetic code, the canonical genetic code being used in early-branching Ulvophyceae and the Bryopsidales, and the alternative code being used in Dasycladales, Cladophorales and Trentepohliales (Fig. 6). Several explanations for the observation that the Bryopsidales, which have the canonical code, are nested in a clade of which all other representatives have the alternative code are compared by Cocquyt et al. (2010a). These authors suggest that a stepwise acquisition model involving ambiguous intermediates (Fig. 6) (Schultz & Yarus, 1994; Santos et al., 2004) could have caused this pattern, but this hypothesis awaits confirmation from studies of tRNA populations and eukaryotic release factor sequences.

The elongation factor 1 alpha gene (EF-1a), which has a function in delivering aminoacyl-tRNAs to the eukaryotic ribosome, is a crucial protein in the translation process. Despite this crucial function, not all eukaryotes have this protein, and it has been shown that an elongation factor-like protein (EF-like) can substitute for EF-1α in these taxa (Keeling & Inagaki, 2004). Interestingly, most eukaryote genomes contain either EF-1α or EF-like, although a few have both (Kamikawa et al., 2008). In green algae, these elongation factors show a mutually exclusive but scattered distribution (Noble et al., 2007; Cocquyt et al., 2009; Gile et al., 2009). Whereas the streptophytes have EF-1α, with the exception of Mesostigma, the chlorophytes encode EF-like with the exception of certain ulvophytes. Like the prasinophytes and most core chlorophytes, the early-branching ulvophytes have the EF-like gene whereas the clade comprising Ignatius, the Trentepohliales, Dasycladales, Bryopsidales and Cladophorales have EF-1α (Fig. 6). Regrettably, phylogeny-explicit models of gene gain and loss were unable to determine the presence of either one or both of the genes in the genome of the ancestor of the green lineage and the Ulvophyceae (Cocquyt et al., 2009). Nevertheless, the alternative elongation factors used in different lineages of the green seaweeds can be taken to mark a second considerable alteration of the translation pathway that occurred during the evolution of the Ulvophyceae.
In addition to the changes in genetic code and elongation factors mentioned above, two other sources of evidence point to changes in the translational apparatus of the Ulvophyceae (Cocquyt, 2009). First, patterns of codon usage of eight nuclear housekeeping genes differ considerably between the prasinophytes and Ulvophyceae. Second, the rates of molecular evolution of the 18S ribosomal RNA molecule are considerably elevated in the ulvophycean orders Cladophorales, Dasycladales and Bryopsidales. All this evidence taken together points to profound changes in the translational apparatus of the Ulvophyceae, but this hypothesis awaits confirmation from more detailed studies of the translational pathway.
E. Molecular evolution in the Streptophyta and the origin of land plants
The origin of embryophytic land plants from a green algal ancestor was a major event in the history of life, which influenced the establishment of the entire terrestrial ecosystem and had far-reaching effects on atmospheric chemistry and climate (Berner, 1997; Kenrick & Crane, 1997; Steemans et al., 2009). As noted above (Section II.B.4), although the Streptophyta are not particularly diverse in terms of number of described species, they represent the full range of structural diversity, from scaly unicellular flagellates through unbranched and branched filaments, to the complex three-dimensionally organized tissues of plants. There is little doubt, however, of the phylogenetic placement of embryophytes among the other streptophytes. The structurally simple streptophytes retain a number of ancestral features that can shed light on the factors that led to the success of embryophytes in the terrestrial environment.
The colonization of dry land involved adaptation to new and harsh environmental conditions such as desiccation, temperature fluctuations, the need for support in the absence of a buoyant medium, and UV radiation (Raven, 2000; Waters, 2003; Floyd & Bowman, 2007; Lang et al., 2008). Ecophysiological adaptations of the first land plants included enhanced osmoregulation and osmoprotection, desiccation and freezing tolerance, and heat resistance (Rensing et al., 2008). Several biochemical innovations have been identified, including synthesis and accumulation of protective “sunscreens,” plant growth hormones, isoprene, phenolics, heat shock proteins, and enhanced DNA repair mechanisms (Waters, 2003; Rensing et al., 2008). In addition, several morphological innovations are believed to have allowed successful adaption to life on land and radiation into new niches (Graham et al., 2000) . Some of these are found in one or more algal relatives of embryophytes, including production of extracellular matrices such as sporopollenin and perhaps lignin, the development of differentiated cells and tissues, dorsiventral development (which is important in the development of structures such as leaves), the establishment of intercellular communication networks (plasmodesmata, plant hormones, receptors and their ligands), and the perception of environmental cues (light and gravity), while others, such as life cycle involving alternation of two distinct multicellular generations (a diploid sporophyte and haploid gametophyte), protected embryos, and gas-filled spaces within the plant body appear to be unique to embryophytes (Delwiche et al., 1989; Graham et al., 2000; Bowman et al., 2007; Ligrone et al., 2008). The success of land plants has further been linked to symbiotic associations with mycorrhizal fungi (Simon et al., 1993; Heckman et al., 2001; McCourt et al., 2004).
Important evolutionary transitions and adaptive radiations, such as the origin and diversification of land plants, have been associated with gene family expansions resulting from large-scale gene duplication or whole-genome duplication events (Ohno, 1970; De Bodt et al., 2005; Flagel & Wendel, 2009). The various physiological and morphological adaptations to land were likely associated with expansion of gene families involved in signalling pathways, such as those for auxin, ABA and cytokinin (Lang et al., 2008; Rensing et al., 2008; Timme & Delwiche, 2010; De Smet et al., 2011). Morphological innovations and the evolution of morphological complexity in land plants have been linked with increased gene family complexity of several genes including actin (An et al., 1999), MADS box genes (Henschel et al., 2002; Tanabe et al., 2005), homeobox genes (Mukherjee et al., 2009) and OPR genes (Li et al., 2009). Expansion of the glutaredoxins gene family (enzymes implicated in oxidative stress response) in land plants likely resulted in genes with novel functions in development and pathogenesis response (Ziemann et al., 2009). The unique sexual life cycle of land plants likely evolved through expansion of homeodomain gene networks (e.g. MADS-box genes) (Tanabe et al., 2005; Niklas & Kutschera, 2010). Ca2+-dependent signalling processes, which are important in the response to many developmental and environmental stimuli, have been found to be very different in green algae and land plants, with several Ca2+ signalling mechanisms have apparently been lost in land plants after their divergence from charophyte algae (Wheeler & Brownlee, 2008).
Although there is still uncertainty concerning the precise relationships between land plants and their green algal relatives (see section II.B.4) it is clear that several fundamental land plant features were inherited by their charophyte green algal progenitors, including cellulosic cell walls, multicellularity, cytokinetic phragmoplast, plasmodesmata, apical meristematic cells (although the meristematic regions in the Charophyceae, Coleochaetophyceae and Zygnematophyceae are very different from that in the embryophytes), asymmetrical cell division, cell specialisation, branching, three dimensional organisation, zygote retention, and placenta (Graham et al., 2000). It has been hypothesized that charophyte green algae were physiologically pre-adapted to life on land by their primary freshwater life style, which exposed them to periodic exposure and desiccation, and allowed a gradual shift towards moist terrestrial habitats, and ultimately the colonization of dry land (Becker & Marin, 2009). It is striking that several streptophyte lineages other than embryophytes are found in terrestrial habitats (i.e., Chlorokybus atmosphyticus, many species of Klebsormidium, and several Zygnematales), which emphasizes the close relationship between freshwater and moist terrestrial habitats.
A growing body of evidence indicates that the molecular bases of many morphological, life-history, cytological, biochemical and physiological features of embryophytes lie in an era prior to the colonization of land (Becker & Marin, 2009). This has been shown by several studies elucidating the evolution of land plant genes in the green lineage, including actin (An et al., 1999), cellulose synthase (Roberts & Roberts, 2004; Yin et al., 2009), class III homeodomain–leucine zipper genes (Floyd et al., 2006), MADS-box genes (Tanabe et al., 2005), vacuolar sorting receptors (Becker & Hoef-Emden, 2009), arabinogalactan-like proteins and hemicelluloses (e.g. endotransglucosylase, responsible for cell wall loosening and cell expansion) (Van Sandt et al., 2007; Eder et al., 2008; Fry et al., 2008).
Expressed sequence tags (ESTs) sequencing of various charophytes and land plants have been used in comparative plant genomics studies to uncover the origins of land plant genes and their associated molecular pathways (Nedelcu et al., 2006; Simon et al., 2006; Timme & Delwiche, 2010). Overall, these studies indicate that several land plant specific characteristics evolved before the transition to land. For example, EST analysis of Mesostigma (Mesostigmatophyceae) suggests that important physiological changes involving regulation of photosynthesis and photorespiration took place early in the evolution of the Streptophyta (Simon et al., 2006). Similarly, putative homologs of genes involved in the development of three-dimensional tissues through asymmetrically dividing apical meristematic cells (BIB gene family) have been identified in Mesostigma (Nedelcu et al., 2006). Several other genes that have been hypothesized to be important in the colonization of land plants (Graham et al., 2000) may have true orthologs in Coleochaete (Coleochaetophyceae) and Spirogyra (Zygnematophyceae) but are apparently absent in the ESTs of the earlier diverging Mesostigma (Timme & Delwiche, 2010). These include genes involved in cellulose synthesis (RSW1), phragmoplast mediated cytokinetic (GEM1/MOR1), formation of plasmodesmata (CRT1) and development of a multicellular sporophyte (MERISTEM LAYER1). Two genes associated with asymmetric cell division (WUSCHEL and GNOM) were only found in Coleochaete, while the plant cell wall loosening expansin genes (EXP) were only present in Spirogyra. In addition, several ethylene pathway genes, long thought to be unique to land plants, have been identified in Coleochaete and Spirogyra.
Cell walls are believed to have played crucial roles in the colonization of land by plants (Sorensen et al., 2010). Although some cell wall components do appear to be land plant innovations, cell wall evolution after the colonization of land appears to be characterized mostly by the elaboration of a pre-existing set of cell wall polysaccharides and the enzymes that synthesize them (e.g. cellulose synthase and wall-remodelling enzymes), rather than substantial innovation (Roberts & Roberts, 2004; Domozych et al., 2007; Van Sandt et al., 2007; Eder et al., 2008; Fry et al., 2008; Domozych et al., 2009; Yin et al., 2009; Popper & Tuohy, 2010; Popper et al., 2011) .
There are several other examples where the genetic potential for plant specific features has been suggested in green algae. For example, genes involved in auxin signalling (central to land plant growth and development) have been detected in various members of Chlorophyta and Streptophyta, indicating that auxin response and transport mechanisms were likely present before the evolution of land plants (De Smet et al., 2011). Similarly, the B3 DNA Binding superfamily, including genes mainly involved in hormone signaling pathways such as those for auxin, abscisic acid, brassinosteroid and gibberellins, are also present in Chlorophyta but have undergone extensive duplication events during land plant evolution (Romanel et al., 2009).
V. Conclusions and perspectives
20 years ago, a review article in this journal reported on the state of knowledge on green algal relationships, gathered from the few pioneering years of ribosomal DNA-based phylogenetic research (Chapman et al., 1991). The past two decades have witnessed profound changes in our understanding of the evolution of green algae. 18S phylogenies have made way for multi-gene and genome scale analyses. These studies have greatly improved our understanding of the deepest relationships of the green lineage (Fig. 3); however, many questions remain.
A large body of molecular evidence has confirmed the ultrastructural-based hypothesis that the green lineage diverged into two discrete clades: the Chlorophyta, which includes the majority of described species of green algae, and the Streptophyta, which is comprised of the charophytes, a paraphyletic assemblage of freshwater algae from which the land plants have evolved. The prasinophytes take up a critical position, diverging early from the remaining Chlorophyta, but the relationships among these lineages remain largely unresolved, mainly because multi-gene data are only available for a limited number of taxa. Similarly, phylogenetic relationships among and within the main clades of the core chlorophytes (Ulvophyceae, Trebouxiophyceae and Chlorophyceae) have not been fully resolved. Other outstanding issues include elucidation of the early diversification of the Streptophyta and identification of the closest living relative of the land plants.
It has become clear that an accurate phylogenetic reconstruction of an ancient group like the green plants will require a rich sampling both in terms of exemplar taxa and molecular markers, along with the application of state of the art phylogenetic techniques. The rapid increase in the amount of genomic data from a wide range of green algae has great potential to resolve large-scale green algal relationships. In addition these data form the basis for investigations of molecular evolution of genes and genomes, providing valuable insight into the evolutionary histories of the green algae.
Acknowledgements
We are grateful to Pavel Skaloud, Fabio Rindi, Juan Lopez-Bautista, Frederick Zechman, Marvin Fawley, Aurora Nedelcu, Marc Buchheim, Andrey Gontcharov, Burkhard Becker and Richard McCourt for discussions and comments on the manuscript. We thank Bob Andersen, Jordi Regas, Antonio Guillén, William Bourland, Jason Oyadomari, Giuseppe Vago, Tom Schils, Antoni López-Arenas, Nadia Abdelahad and Deborah Shelton for providing photographs.
References
(references indicated with * have been updated since Jan 2012)
Adl, S. M., Simpson, A. G. B., Farmer, M. A., Andersen, R. A., Anderson, O. R., et al. 2005. The new higher level classification of eukaryotes with emphasis on the taxonomy of protists. J. Eukaryot. Microbiol. 52: 399-451.
Alberghina, J. S., Vigna, M. S. and Confalonieri, V. A. 2006. Phylogenetic position of the Oedogoniales within the green algae (Chlorophyta) and the evolution of the absolute orientation of the flagellar apparatus. Plant Syst. Evol. 261: 151-163.
An, S. S., Mopps, B., Weber, K. and Bhattacharya, D. 1999. The origin and evolution of green algal and plant actins. Mol. Biol. Evol. 16: 275-285.
Archibald, J. M. 2009. The puzzle of plastid evolution. Curr. Biol. 19: R81-R88.
Aslam, Z., Shin, W. G., Kim, M. K., Im, W. T. and Lee, S. T. 2007. Marinichlorella kaistiae gen. et sp. nov. (Trebouxiophyceae, Chlorophyta) based on polyphasic taxonomy. J. Phycol. 43: 576-584.
Bakker, F. T., Olsen, J. L., Stam, W. T. and Van den Hoek, C. 1994. The Cladophora complex (Chlorophyta): new views based on 18S rRNA gene sequences. Mol. Phylogenet. Evol. 3: 365-382.
Baldauf, S. L. 2008. An overview of the phylogeny and diversity of eukaryotes. J. Syst. Evol. 46: 263-273.
Baldauf, S. L., Manhart, J. R. and Palmer, J. D. 1990. Different fates of the chloroplast tufA gene following its transfer to the nucleus in green algae. Proc. Natl. Acad. Sci. U.S.A. 87: 5317-5321.
Baldauf, S. L. and Palmer, J. D. 1990. Evolutionary transfer of the chloroplast tufA gene to the nucleus. Nature 344: 262-265.
Ballantine, D. L. and Norris, J. N. 1994. Verdigellas, a new deep-water genus (Tetrasporales, Chlorophyta) from the tropical western Atlantic. Crypt. Bot. 4: 368-372.
Baurain, D., Brinkmann, H., Petersen, J., Rodriguez-Ezpeleta, N., Stechmann, A., Demoulin, V., Roger, A. J., Burger, G., Lang, B. F. and Philippe, H. 2010. Phylogenomic evidence for separate acquisition of plastids in cryptophytes, haptophytes, and stramenopiles. Mol. Biol. Evol. 27: 1698-1709.
Barsanti, L., Frassanito, A. M., Passarelli, V., Evangelista, V., Etebari, M., Paccagnini, E., Lupetti, P., Lenzi, P., Verni, F. and Gualtieri, P. 2013. Tetraflagellochloris mauritanica gen. et sp. nov. (Chlorophyceae), a new flagellated alga from the Mauritanian desert: Morphology, ultrastructure, and phylogenetic framing. J. Phycol. 49: 178-193.
Becker, B. and Hoef-Emden, K. 2009. Evolution of vacuolar targeting in algae. Bot. Mar. 52: 117-128.
Becker, B. and Marin, B. 2009. Streptophyte algae and the origin of embryophytes. Annals of Botany 103: 999-1004.
Becker, B., Marin, B. and Melkonian, M. 1994. Structure, composition, and biogenesis of prasinophyte cell coverings. Protoplasma 181: 233-244.
Bélanger, A. S., Brouard, J. S., Charlebois, P., Otis, C., Lemieux, C. and Turmel, M. 2006. Distinctive architecture of the chloroplast genome in the chlorophycean green alga Stigeoclonium helveticum. Mol. Genet. Genomics 276: 464-477.
Bendich, A. J. 2004. Circular chloroplast chromosomes: The grand illusion. Plant Cell 16: 1661-1666.
Bendich, A. J. 2007. The size and form of chromosomes are constant in the nucleus, but highly variable in bacteria, mitochondria and chloroplasts. BioEssays 29: 474-483.
Berger, S., Fettweiss, U., Gleissberg, S., Liddle, L. B., Richter, U., Sawitzky, H. and Zuccarello, G. C. 2003. 18S rDNA phylogeny and evolution of cap development in Polyphysaceae (formerly Acetabulariaceae; Dasycladales, Chlorophyta). Phycologia 42: 506-561.
Berger, S. and Kaever, M. J. 1992. Dasycladales: an illustrated monograph of a fascinating algal order. Thieme, Stuttgart.
Berner, R. A. 1997. Paleoclimate – The rise of plants and their effect on weathering and atmospheric CO2. Science 276: 544-546.
Berney, C. and Pawlowski, J. 2006. A molecular time-scale for eukaryote evolution recalibrated with the continuous microfossil record. Proc. R. Soc. B Biol. Sci. 273: 1867-1872.
Bhattacharya, D., Friedl, T. and Damberger, S. 1996. Nuclear-encoded rDNA group I introns: Origin and phylogenetic relationships of insertion site lineages in the green algae. Mol. Biol. Evol. 13: 978-989.
Bhattacharya, D. and Medlin, L. 1998. Algal phylogeny and the origin of land plants. Plant Physiol. 116: 9-15.
Bhattacharya, D., Weber, K., An, S. S. and Berning-Koch, W. 1998. Actin phylogeny identifies Mesostigma viride as a flagellate ancestor of the land plants. J. Mol. Evol. 47: 544-550.
Bisson, M. A., Beilby, M. J. and Shepherd, V. A. 2006. Electrophysiology of turgor regulation in marine siphonous green algae. J. Membr. Biol. 211: 1-14.
Blackman, F. 1900. The primitive algae and the Flagellata. Annals of Botany 14: 647-688.
Blaha, J., Baloch, E. and Grube, M. 2006. High photobiont diversity associated with the euryoecious lichen-forming ascomycete Lecanora rupicola (Lecanoraceae, Ascomycota). Biol. J. Linn. Soc. 88: 283-293.
Blanc, G., Duncan, G., Agarkova, I., Borodovsky, M., Gurnon, J., et al. 2010. The Chlorella variabilis NC64A genome reveals adaptation to photosymbiosis, coevolution with viruses, and cryptic sex. Plant Cell 22: 2943-2955.
Bodyl, A., Stiller, J. W. and Mackiewicz, P. 2009. Chromalveolate plastids: direct descent or multiple endosymbioses? Trends Ecol. Evol. 24: 119-121.
Boer, P. H. and Gray, M. W. 1986. The URF 5 gene of Chlamydomonas reinhardtii mitochondria: DNA sequence and mode of transcription. EMBO J. 5: 21-28.
Boer, P. H. and Gray, M. W. 1988. Genes encoding a subunit of respiratory NADH dehydrogenase (ND1) and a reverse transcriptase-like protein (RTL) are linked to ribosomal RNA gene pieces in Chlamydomonas reinhardtii mitochondrial DNA. EMBO J. 7: 3501-3508.
Boer, P. H. and Gray, M. W. 1991. Short dispersed repeats localized in spacer regions of Chlamydomonas reinhardtii mitochondrial DNA. Curr. Genet. 19: 309-312.
Booton, G. C., Floyd, G. L. and Fuerst, P. A. 1998a. Origins and affinities of the filamentous green algal orders Chaetophorales and Oedogoniales based on 18S rRNA gene sequences. J. Phycol. 34: 312-318.
Booton, G. C., Floyd, G. L. and Fuerst, P. A. 1998b. Polyphyly of tetrasporalean green algae inferred from nuclear small-subunit ribosomal DNA. J. Phycol. 34: 306-311.
Borza, T., Redmond, E. K., Laflamme, M. and Lee, R. W. 2009. Mitochondrial DNA in the Oogamochlamys clade (Chlorophyceae): high GC content and unique genome architecture for green algae. J. Phycol. 45: 1323-1334.
Boudreau, E., Otis, C. and Turmel, M. 1994. Conserved gene clusters in the highly rearranged chloroplast genomes of Chlamydomonas moewusii and Chlamydomonas reinhardtii. Plant Mol. Biol. 24: 585-602.
Boudreau, E. and Turmel, M. 1996. Extensive gene rearrangements in the chloroplast DNAs of Chlamydomonas species featuring multiple dispersed repeats. Mol. Biol. Evol. 13: 233-243.
Bowman, J. L., Floyd, S. K. and Sakakibara, K. 2007. Green genes – Comparative genomics of the green branch of life. Cell 129: 229-234.
Bremer, K. 1985. Summary of green plant phylogeny and classification. Cladistics 1: 369-385.
Breton, G. and Kay, S. 2006. Circadian rhythms lit up in Chlamydomonas. Genome Biology 7: 215.
Brouard, J.-S., Otis, C., Lemieux, C. and Turmel, M. 2008. Chloroplast DNA sequence of the green alga Oedogonium cardiacum (Chlorophyceae): Unique genome architecture, derived characters shared with the Chaetophorales and novel genes acquired through horizontal transfer. BMC Genomics 9: 290.
Brouard, J.-S., Otis, C., Lemieux, C. and Turmel, M. 2010. The exceptionally large chloroplast genome of the green alga Floydiella terrestris illuminates the evolutionary history of the Chlorophyceae. Genome Biol. Evol. 2: 240-256.
Brouard, J.-S., Otis, C., Lemieux, C. and Turmel, M. 2011. The chloroplast genome of the green alga Schizomeris leibleinii (Chlorophyceae) provides evidence for bidirectional DNA replication from a single origin in the Chaetophorales. Genome Biol. Evol.: doi:10.1093/gbe/evr037.
Buchheim, M., Buchheim, J., Carlson, T., Braband, A., Hepperle, D., Krienitz, L., Wolf, M. and Hegewald, E. 2005. Phylogeny of the Hydrodictyaceae (Chlorophyceae): Inferences from rDNA data. J. Phycol. 41: 1039-1054.
Buchheim, M. A. and Buchheim, J. A. 2001. Phylogeny of Geminella (Chlorophyta) and allies: a study of 18S rDNA sequences. http://www.bio.utulsa.edu/deepestgreen/Geminella.htm.
Buchheim, M. A. and Chapman, R. L. 1991. Phylogeny of the colonial green flagellates: a study of 18S and 26S rRNA sequence data. Biosystems 25: 85-100.
Buchheim, M. A., Kirkwood, A. E., Buchheim, J. A., Verghese, B. and Henley, W. J. 2010. Hypersaline soil supports a diverse community of Dunaliella (Chlorophyceae). J. Phycol. 46: 1038-1047.
Buchheim, M. A., Lemieux, C., Otis, C., Gutell, R. R., Chapman, R. L. and Turmel, M. 1996. Phylogeny of the Chlamydomonadales (Chlorophyceae): A comparison of ribosomal RNA gene sequences from the nucleus and the chloroplast. Mol. Phylogenet. Evol. 5: 391-402.
Buchheim, M. A., McAuley, M. A., Zimmer, E. A., Theriot, E. C. and Chapman, R. L. 1994. Multiple origins of colonial green flagellates from unicells: Evidence from molecular and organismal characters. Mol. Phylogenet. Evol. 3: 322-343.
Buchheim, M. A., Michalopulos, E. A. and Buchheim, J. A. 2001. Phylogeny of the Chlorophyceae with special reference to the Sphaeropleales: A study of 18S and 26S rDNA data. J. Phycol. 37: 819-835.
Buchheim, M. A., Turmel, M., Zimmer, E. A. and Chapman, R. L. 1990. Phylogeny of Chlamydomonas (Chlorophyta) based on cladistic analysis of nuclear 18S rRNA sequence data. J. Phycol. 26: 689-699.
Butcher, R. W. 1952. Contribution to our knowledge of the smaller marine algae. J. Mar. Biol. Assoc. U. K. 31: 175-191.
Butterfield, N. J. 2009. Modes of pre-Ediacaran multicellularity. Precambrian Res. 173: 201-211.
Butterfield, N. J., Knoll, A. H. and Swett, K. 1994. Paleobiology of the Neoproterozoic Svanbergfjellet Formation, Spitsbergen. Fossils and Strata 34: 1-84.
Cachon, M. and Caram, B. 1979. A symbiotic green alga, Pedinomonas symbiotica sp. nov. (Prasinophyceae), in the radiolarian Thalassolampe margarodes. Phycologia 18: 177-184.
Caisová, L., Marin, B., Sausen, N., Pröschold, T. and Melkonian, M. 2011. Polyphyly of Chaetophora and Stigeoclonium within the Chaetophorales (Chlorophyceae), revealed by sequence comparisons of nuclear-encoded SSU rRNA genes. J. Phycol. 47: 164-177.
Calvin, M. and Benson, A. A. 1948. The path of carbon in photosynthesis. Science 107: 476-480.
Cardol, P., Bailleul, B., Rappaport, F., Derelle, E., Beal, D., Breyton, C., Bailey, S., Wollman, F. A., Grossman, A., Moreau, H. and Finazzi, G. 2008. An original adaptation of photosynthesis in the marine green alga Ostreococcus. Proc. Natl. Acad. Sci. U.S.A. 105: 7881-7886.
Cavalier-Smith, T. 1981. Eukaryote kingdoms: seven or nine? Biosyst. Eng. 14: 461-481.
Cavalier-Smith, T. 1998. A revised six-kingdom system of life. Biol. Rev. Camb. Philos. Soc. 73: 203-266.
Cavalier-Smith, T. 1999. Principles of protein and lipid targeting in secondary symbiogenesis: Euglenoid, dinoflagellate, and sporozoan plastid origins and the eukaryote family tree. J. Eukaryot. Microbiol. 46: 347-366.
Cavalier-Smith, T. 2002. Nucleomorphs: enslaved algal nuclei. Curr. Opin. Microbiol. 5: 612-619.
Cavalier-Smith, T. 2006. Cell evolution and earth history: stasis and revolution. Philos. Trans. R. Soc. Lond. B Biol. Sci. 361: 969-1006.
Chapman, R. L. 1984. An assessment of the current state of our knowledge of the Trentepohliaceae. In: Systematics of the green algae. pp. 233-250. Irvine, D. E. G. & John, D. M. Eds., Academic Press, London.
Chapman, R. L., Borkhsenious, O. L., Brown, R. C., Henk, M. C. and Waters, D. A. 2001. Phragmoplast-mediated cytokinesis in Trentepohlia: results of TEM and immunofluorescence cytochemistry. Int. J. Syst. Evol. Microbiol. 51: 759-765.
Chapman, R. L., Buchheim, M. A., Delwiche, C. F., Friedl, T., Huss, V. A. R., Karol, K. G., Lewis, L. A., Manhart, J., McCourt, R. M., Olsen, J. L. and Waters, D. A. 1998. Molecular systematics of the green algae. In: Molecular Systematics of Plants II. pp. 508-540. Soltis, D. E., Soltis, P. S. & Doyle, J. J. Eds., Kluwer Academic Publishers, Boston.
Chapman, R. L., Buchheim, M. A. and Hoshaw, R. W. 1991. Ribosomal RNA gene sequences: Analysis and significance in the phylogeny and taxonomy of green algae. Crit. Rev. Plant Sci. 10: 343-368.
Chappell, D. F., Okelly, C. J. and Floyd, G. L. 1991. Flagellar apparatus of the biflagellate zoospores of the enigmatic marine green-alga Blastophysa rhizopus. J. Phycol. 27: 423-428.
Cheng, Q., Fowler, R., Tam, L. W., Edwards, L. and Miller, S. M. 2003. The role of GlsA in the evolution of asymmetric cell division in the green alga Volvox carteri. Dev. Genes Evol. 213: 328-335.
Cheung, M. K., Au, C. H., Chu, K. H., Kwan, H. S. and Wong, C. K. 2010. Composition and genetic diversity of picoeukaryotes in subtropical coastal waters as revealed by 454 pyrosequencing. ISME Journal 4: 1053-1059.
Chihara, M., Inouye, I. and Takahata, N. 1986. Oltmannsiellopsis, a new genus of marine flagellate (Dunaliellaceae, Chlorophyceae). Arch. Protistenk. 132: 313-324.
Chisholm, J. R. M., Dauga, C., Ageron, E., Grimont, P. A. D. and Jaubert, J. M. 1996. ‘Roots’ in mixotrophic algae. Nature 381: 382-382.
Cocquyt, E. 2009. Phylogeny and molecular evolution of green algae. PhD, Ghent University, 167 pp.
Cocquyt, E., Gile, G., Leliaert, F., Verbruggen, H., Keeling, P. and De Clerck, O. 2010a. Complex phylogenetic distribution of a non-canonical genetic code in green algae. BMC Evol. Biol. 10: 327.
Cocquyt, E., Verbruggen, H., Leliaert, F. and De Clerck, O. 2010b. Evolution and cytological diversification of the green seaweeds (Ulvophyceae). Mol. Biol. Evol. 27: 2052-2061.
Cocquyt, E., Verbruggen, H., Leliaert, F., Zechman, F., Sabbe, K. and De Clerck, O. 2009. Gain and loss of elongation factor genes in green algae. BMC Evol. Biol. 9: 39.
Colbath, G. K. 1983. Fossil prasinophycean phycomata (Chlorophyta) from the Silurian Bainbridge formation, Missouri, USA. Phycologia 22: 249-265.
Cook, M. E. 2004a. Cytokinesis in Coleochaete orbicularis (Charophyceae): An ancestral mechanism inherited by plants. Am. J. Bot. 91: 313-320.
Cook, M. E. 2004b. Structure and asexual reproduction of the enigmatic charophycean green alga Entransia fimbriata (Klebsormidiales, Charophyceae). J. Phycol. 40: 424-431.
Countway, P. D. and Caron, D. A. 2006. Abundance and distribution of Ostreococcus sp. in the San Pedro Channel, California, as revealed by quantitative PCR. Appl. Environ. Microbiol. 72: 2496-2506.
Courties, C., Vaquer, A., Troussellier, M., Lautier, J., Chretiennot-Dinet, M. J., Neveux, J., Machado, C. and Claustre, H. 1994. Smallest eukaryotic organism. Nature 370: 255-255.
Dagan, T. and Martin, W. 2009a. Getting a better picture of microbial evolution en route to a network of genomes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 364: 2187-2196.
Dagan, T. and Martin, W. 2009b. Seeing green and red in diatom genomes. Science 324: 1651-1652.
Daugbjerg, N., Moestrup, Ø. and Arctander, P. 1994. Phylogeny of the genus Pyramimonas (Prasinophyceae, Chlorophyta) inferred from the rbcL gene. J. Phycol. 30: 991-999.
Daugbjerg, N., Moestrup, Ø. and Arctander, P. 1995. Phylogeny of genera of Prasinophyceae and Pedinophyceae (Chlorophyta) deduced from molecular analysis of the rbcL gene. Phycol. Res. 43: 203-213.
De Bodt, S., Maere, S. and Van de Peer, Y. 2005. Genome duplication and the origin of angiosperms. Trends Ecol. Evol. 20: 591-597.
de Cambiaire, J. C., Otis, C., Lemieux, C. and Turmel, M. 2006. The complete chloroplast genome sequence of the chlorophycean green alga Scenedesmus obliquus reveals a compact gene organization and a biased distribution of genes on the two DNA strands. BMC Evol. Biol. 6: 37.
de Cambiaire, J. C., Otis, C., Turmel, M. and Lemieux, C. 2007. The chloroplast genome sequence of the green alga Leptosira terrestris: multiple losses of the inverted repeat and extensive genome rearrangements within the Trebouxiophyceae. BMC Genomics 8: 213.
de Jesus, M. D., Tabatabai, F. and Chapman, D. J. 1989. Taxonomic distribution of copper-zinc superoxide dismutase in green algae and its phylogenetic importance. J. Phycol. 25: 767-772.
de Koning, A. P. and Keeling, P. J. 2004. Nucleus-encoded genes for plastid-targeted proteins in Helicosporidium: Functional diversity of a cryptic plastid in a parasitic alga. Eukaryotic Cell 3: 1198-1205.
de Koning, A. P. and Keeling, P. J. 2006. The complete plastid genome sequence of the parasitic green alga Helicosporidium sp. is highly reduced and structured. BMC Biology 4: 12.
de Koning, A. P., Tartar, A., Boucias, D. G. and Keeling, P. J. 2005. Expressed sequence tag (EST) survey of the highly adapted green algal parasite, Helicosporidium. Protist 156: 181-190.
De Smet, I., Voss, U., Lau, S., Wilson, M., Shao, N., Timme, R. E., Swarup, R., Kerr, I., Hodgman, C., Bock, R., Bennett, M., Jurgens, G. and Beeckman, T. 2011. Unraveling the evolution of auxin signaling. Plant Physiol. 155: 209-221.
De Wever, A., Leliaert, F., Verleyen, E., Vanormelingen, P., Van der Gucht, K., Hodgson, D. A., Sabbe, K. and Vyverman, W. 2009. Hidden levels of phylodiversity in Antarctic green algae: further evidence for the existence of glacial refugia. Proc. R. Soc. B Biol. Sci. 276: 3591-3599.
Deason, T. R., Silva, P. C., Watanabe, S. and Floyd, G. L. 1991. Taxonomic status of the species of the green algal genus Neochloris. Plant Syst. Evol. 177: 213-219.
Delwiche, C. F. 1999. Tracing the thread of plastid diversity through the tapestry of life. Am. Nat. 154: S164-S177.
Delwiche, C. F., Graham, L. E. and Thomson, N. 1989. Lignin-like compounds and sporopollenin in Coleochaete, an algal model for land plant ancestry. Science 245: 399-401.
Delwiche, C. F., Karol, K. G., Cimino, M. T. and Sytsma, K. J. 2002. Phylogeny of the genus Coleochaete (Coleochaetales, Charophyta) and related taxa inferred by analysis of the chloroplast gene rbcL. J. Phycol. 38: 394-403.
Demir-Hilton, E., Sudek, S., Cuvelier, M. L., Gentemann, C. L., Zehr, J. P. and Worden, A. Z. 2011. Global distribution patterns of distinct clades of the photosynthetic picoeukaryote Ostreococcus. ISME Journal: doi:10.1038/ismej.2010.209.
Denboh, T., Hendrayanti, D. and Ichimura, T. 2001. Monophyly of the genus Closterium and the order Desmidiales (Charophyceae, Chlorophyta) inferred from nuclear small subunit rDNA data. J. Phycol. 37: 1063-1072.
Derelle, E., Ferraz, C., Rombauts, S., Rouze, P., Worden, A. Z., et al. 2006. Genome analysis of the smallest free-living eukaryote Ostreococcus tauri unveils many unique features. Proc. Natl. Acad. Sci. U.S.A. 103: 11647-11652.
Diez, B., Pedros-Alio, C. and Massana, R. 2001. Study of genetic diversity of eukaryotic picoplankton in different oceanic regions by small-subunit rRNA gene cloning and sequencing. Appl. Environ. Microbiol. 67: 2932-2941.
Domozych, D. S., Lambiasse, L., Kiemle, S. N. and Gretz, M. R. 2009. Cell-wall development and bipolar growth in the desmid Penium margaritaceum (Zygnematophyceae, Streptophyta). Asymmetry in a symmetric world. J. Phycol. 45: 879-893.
Domozych, D. S., Serfis, A., Kiemle, S. N. and Gretz, M. R. 2007. The structure and biochemistry of charophycean cell walls: I. Pectins of Penium margaritaceum. Protoplasma 230: 99-115.
Doolittle, W. E. 1998. You are what you eat: a gene transfer ratchet could account for bacterial genes in eukaryotic nuclear genomes. Trends Genet. 14: 307-311.
Douzery, E. J. P., Snell, E. A., Bapteste, E., Delsuc, F. and Philippe, H. 2004. The timing of eukaryotic evolution: Does a relaxed molecular clock reconcile proteins and fossils? Proc. Natl. Acad. Sci. U.S.A. 101: 15386-15391.
Drummond, C. S., Hall, J., Karol, K. G., Delwiche, C. F. and McCourt, R. M. 2005. Phylogeny of Spirogyra and Sirogonium (Zygnematophyceae) based on rbcL sequence data. J. Phycol. 41: 1055-1064.
Duncan, L., Nishii, I., Harryman, A., Buckley, S., Howard, A., Friedman, N. R. and Miller, S. M. 2007. The VARL gene family and the evolutionary origins of the master cell-type regulatory gene, regA, in Volvox carteri. J. Mol. Evol. 65: 1-11.
Dupuy, L., Mackenzie, J. and Haseloff, J. 2010. Coordination of plant cell division and expansion in a simple morphogenetic system. Proc. Natl. Acad. Sci. U.S.A. 107: 2711-2716.
Eddie, B., Krembs, C. and Neuer, S. 2008. Characterization and growth response to temperature and salinity of psychrophilic, halotolerant Chlamydomonas sp. ARC isolated from Chukchi Sea ice. Mar. Ecol. Prog. Ser. 354: 107-117.
Eder, M., Tenhaken, R., Driouich, A. and Lütz-Meindl, U. 2008. Occurrence and characterization of arabinogalactan-like proteins and hemicelluloses in Micrasterias (Streptophyta). J. Phycol. 44: 1221-1234.
Edgcomb, V. P., Kysela, D. T., Teske, A., Gomez, A. D. and Sogin, M. L. 2002. Benthic eukaryotic diversity in the Guaymas Basin hydrothermal vent environment. Proc. Natl. Acad. Sci. U.S.A. 99: 7658-7662.
Eikrem, W. and Throndsen, J. 1990. The ultrastructure of Bathycoccus gen. nov. and Bathycoccus prasinos sp. nov., a nonmotile picoplanktonic alga (Chlorophyta, Prasinophyceae) from the Mediterranean and Atlantic. Phycologia 29: 344-350.
Elias, M. and Archibald, J. M. 2009. Sizing up the genomic footprint of endosymbiosis. BioEssays 31: 1273-1279.
Elias, M. and Neustupa, J. 2009. Pseudomarvania, gen. nov. (Chlorophyta, Trebouxiophyceae), a new genus for “budding” subaerial green algae Marvania aerophytica Neustupa et Šejnohová and Stichococcus ampulliformis Handa. Fottea 9: 169-178.
Falkowski, P. G., Katz, M. E., Knoll, A. H., Quigg, A., Raven, J. A., Schofield, O. and Taylor, F. J. R. 2004. The evolution of modern eukaryotic phytoplankton. Science 305: 354-360.
Fan, J. and Lee, R. W. 2002. Mitochondrial genome of the colorless green alga Polytomella parva: Two linear DNA molecules with homologous inverted repeat termini. Mol. Biol. Evol. 19: 999-1007.
Fawley, M. W., Yun, Y. and Qin, M. 2000. Phylogenetic analyses of 18S rDNA sequences reveal a new coccoid lineage of the Prasinophyceae (Chlorophyta). J. Phycol. 36: 387-393.
Feist, M. and Feist, R. 1997. Oldest record of a bisexual plant. Nature 385: 401-401.
Ferris, P., Olson, B., De Hoff, P. L., Douglass, S., Casero, D., et al. 2010. Evolution of an expanded sex-determining locus in Volvox. Science 328: 351-354.
Ferris, P. J., Armbrust, E. V. and Goodenough, U. W. 2002. Genetic structure of the mating-type locus of Chlamydomonas reinhardtii. Genetics 160: 181-200.
Ferris, P. J. and Goodenough, U. W. 1994. The mating-type locus of Chlamydomonas reinhardtii contains highly rearranged DNA sequences. Cell 76: 1135-1145.
Ferris, P. J. and Goodenough, U. W. 1997. Mating type in Chlamydomonas is specified by mid, the minus-dominance gene. Genetics 146: 859-869.
Finet, C., Timme, R. E., Delwiche, C. F. and Marlétaz, F. 2010. Multigene phylogeny of the green lineage reveals the origin and diversification of land plants. Curr. Biol. 20: 2217-2222.
* Finet, C., Timme, R. E., Delwiche, C. F. and Marlétaz, F. 2012. Erratum – Multigene phylogeny of the green lineage reveals the origin and diversification of land plants. Curr. Biol. 22: 1456-1457.
Flagel, L. E. and Wendel, J. F. 2009. Gene duplication and evolutionary novelty in plants. New Phytol. 183: 557-564.
Floyd, G. L., Hoops, H. J. and Swanson, J. A. 1980. Fine structure of the zoospore of Ulothrix belkae with emphasis on the flagellar apparatus. Protoplasma 104: 17-31.
Floyd, S. K. and Bowman, J. L. 2007. The ancestral developmental tool kit of land plants. Int. J. Plant Sci. 168: 1-35.
Floyd, S. K., Zalewski, C. S. and Bowman, J. L. 2006. Evolution of class III homeodomain-leucine zipper genes in streptophytes. Genetics 173: 373-388.
Fott, B. 1971. Algenkunde, 2nd Ed. VEB Fischer, Jena.
Foulon, E., Not, F., Jalabert, F., Cariou, T., Massana, R. and Simon, N. 2008. Ecological niche partitioning in the picoplanktonic green alga Micromonas pusilla: evidence from environmental surveys using phylogenetic probes. Environ. Microbiol. 10: 2433-2443.
Frederick, S. E., Gruber, P. J. and Tolbert, N. E. 1973. The occurrence of glycolate dehydrogenase and glycolate oxidase in green plants: An evolutionary survey. Plant Physiol. 52: 318-323.
Friedl, T. 1995. Inferring taxonomic positions and testing genus level assignments in coccoid green lichen algae: a phylogenetic analysis of 18S ribosomal RNA sequences from Dictyochloropsis reticulata and from members of the genus Myrmecia (Chlorophyta, Trebouxiophyceae cl. nov.). J. Phycol. 31: 632-639.
Friedl, T. 1996. Evolution of the polyphyletic genus Pleurastrum (Chlorophyta): inferences from nuclear-encoded ribosomal DNA sequences and motile cell ultrastructure. Phycologia 35: 456-469.
Friedl, T., Besendahl, A., Pfeiffer, P. and Bhattacharya, D. 2000. The distribution of group I introns in lichen algae suggests that lichenization facilitates intron lateral transfer. Mol. Phylogenet. Evol. 14: 342-352.
Friedl, T. and Bhattacharya, D. 2002. Origin and evolution of green lichen algae. In: Symbiosis. pp. 341-357. Seckbach, J. Ed., Kluwer Academic Publishers, Dordrecht.
Friedl, T. and Büdel, B. 1996. Photobionts. In: Lichen biology. pp. 8-23. Nash, T. H. Ed., Cambridge University Press, Cambridge.
Friedl, T. and O’Kelly, C. J. 2002. Phylogenetic relationships of green algae assigned to the genus Planophila (Chlorophyta): evidence from 18S rDNA sequence data and ultrastructure. Eur. J. Phycol. 37: 373-384.
Fritsch, F. E. 1935. The Structure and Reproduction of the Algae, Vol. I. Cambridge University Press, London.
Fry, S. C., Mohler, K. E., Nesselrode, B. H. W. A. and Franková, L. 2008. Mixed-linkage β-glucan: xyloglucan endotransglucosylase, a novel wall-remodelling enzyme from Equisetum (horsetails) and charophytic algae. The Plant Journal 55: 240-252.
Fulton, A. B. 1978. Colonial development in Pandorina morum.1. Structure and composition of extracellular matrix. Dev. Biol. 64: 224-235.
Gerisch, G. 1959. Die Zelldifferenzierung bei Pleodorina californica Shaw und die Organisation der Phytomonadinenkolonien. Arch. Protistenk. 104: 292-358.
Ghoshroy, S., Binder, M., Tartar, A. and Robertson, D. L. 2010. Molecular evolution of glutamine synthetase II: Phylogenetic evidence of a non-endosymbiotic gene transfer event early in plant evolution. BMC Evol. Biol. 10: 198.
Gile, G. H., Novis, P. M., Cragg, D. S., Zuccarello, G. C. and Keeling, P. J. 2009. The distribution of Elongation Factor-1 alpha (EF-1α), Elongation Factor-Like (EFL), and a non-canonical genetic code in the Ulvophyceae: discrete genetic characters support a consistent phylogenetic framework. J. Eukaryot. Microbiol. 56: 367-372.
Glanz, S. and Kuck, U. 2009. Trans-splicing of organelle introns – a detour to continuous RNAs. BioEssays 31: 921-934.
Gockel, G. and Hachtel, W. 2000. Complete gene map of the plastid genome of the nonphotosynthetic euglenoid flagellate Astasia longa. Protist 151: 347-351.
Gontcharov, A. A. 2008. Phylogeny and classification of Zygnematophyceae (Streptophyta): current state of affairs. Fottea 8: 87-104.
Gontcharov, A. A., Marin, B. and Melkonian, M. 2003. Molecular phylogeny of conjugating green algae (Zygnemophyceae, Streptophyta) inferred from SSU rDNA sequence comparisons. J. Mol. Evol. 56: 89-104.
Gontcharov, A. A., Marin, B. and Melkonian, M. 2004. Are combined analyses better than single gene phylogenies? A case study using SSU rDNA and rbcL sequence comparisons in the Zygnematophyceae (Streptophyta). Mol. Biol. Evol. 21: 612-624.
Gontcharov, A. A. and Melkonian, M. 2004. Unusual position of the genus Spirotaenia (Zygnematophyceae) among streptophytes revealed by SSU rDNA and rbcL sequence comparisons. Phycologia 43: 105-113.
Gontcharov, A. A. and Melkonian, M. 2005. Molecular phylogeny of Staurastrum Meyen ex Ralfs and related genera (Zygnematophyceae, Streptophyta) based on coding and noncoding rDNA sequence comparisons. J. Phycol. 41: 887-899.
Gontcharov, A. A. and Melkonian, M. 2008. In search of monophyletic taxa in the family Desmidiaceae (Zygnematophyceae, Viridiplantae): The genus Cosmarium. Am. J. Bot. 95: 1079-1095.
Gontcharov, A. A. and Melkonian, M. 2010. Molecular phylogeny and revision of the genus Netrium (Zygnematophyceae, Streptophyta): Nucleotaenium gen. nov. J. Phycol. 46: 346-362.
Gontcharov, A. A. and Melkonian, M. 2011. A study of conflict between molecular phylogeny and taxonomy in the Desmidiaceae (Streptophyta, Viridiplantae): analyses of 291 rbcL sequences. Protist 162: 253-267.
Gould, S. B., Waller, R. R. and McFadden, G. I. 2008. Plastid evolution. Annu. Rev. Plant Biol. 59: 491-517.
Gouveia, L. and Oliveira, A. 2009. Microalgae as a raw material for biofuels production. J. Ind. Microbiol. Biotechnol. 36: 269-274.
Graham, L. E. 1982. The occurrence, evolution, and phylogenetic significance of parenchyma in Coleochaete Bréb. (Chlorophyta). Am. J. Bot. 69: 447-454.
Graham, L. E. 1984. Coleochaete and the origin of land plants. Am. J. Bot. 71: 603-608.
Graham, L. E. 1993. Origin of Land Plants. J. Wiley and Sons, New York.
Graham, L. E., Cook, M. E. and Busse, J. S. 2000. The origin of plants: Body plan changes contributing to a major evolutionary radiation. Proc. Natl. Acad. Sci. U.S.A. 97: 4535-4540.
Graham, L. E., Graham, J. M. and Wilcox, L. W. 2009. Algae (2nd edition). Pearson Education, San Francisco.
Graham, L. E. and Kaneko, Y. 1991. Subcellular structures of relevance to the origin of land plants (embryophytes) from green algae. Crit. Rev. Plant Sci. 10: 323-342.
Gray, M. W. 1999. Evolution of organellar genomes. Curr. Opin. Genet. Dev. 9: 678-687.
Gray, M. W. and Boer, P. H. 1988. Organization and expression of algal (Chlamydomonas reinhardtii) mitochondrial DNA. Philos. Trans. R. Soc. Lond. B Biol. Sci. 319: 135-147.
Gray, M. W., Lang, B. F. and Burger, G. 2004. Mitochondria of protists. Annu. Rev. Genet. 38: 477-524.
Gray, M. W. and Schnare, M. N. 1996. Evolution of rRNA gene organization. In: Ribosomal RNA: structure, evolution, recessing and function in protein biosynthesis. pp. 49-69. Zimmerman, R. A. & Dalhberg, A. E. Eds., CRC Press, Boca Raton, Florida.
Green, K. J., Viamontes, G. I. and Kirk, D. L. 1981. Mechanism of formation, ultrastructure and function of the cytoplasmic bridge system during morphogenesis in Volvox. J. Cell Biol. 91: 756-769.
Grimsley, N., Pequin, B., Bachy, C., Moreau, H. and Piganeau, G. 2010. Cryptic sex in the smallest eukaryotic marine green alga. Mol. Biol. Evol. 27: 47-54.
Grossman, A. R., Harris, E. E., Hauser, C., Lefebvre, P. A., Martinez, D., Rokhsar, D., Shrager, J., Silflow, C. D., Stern, D., Vallon, O. and Zhang, Z. D. 2003. Chlamydomonas reinhardtii at the crossroads of genomics. Eukaryotic Cell 2: 1137-1150.
Guillou, L., Eikrem, W., Chretiennot-Dinet, M. J., Le Gall, F., Massana, R., Romari, K., Pedros-Alio, C. and Vaulot, D. 2004. Diversity of picoplanktonic prasinophytes assessed by direct nuclear SSU rDNA sequencing of environmental samples and novel isolates retrieved from oceanic and coastal marine ecosystems. Protist 155: 193-214.
Gunderson, J. H., Elwood, H., Ingold, A., Kindle, K. and Sogin, M. L. 1987. Phylogenetic relationships between chlorophytes, chrysophytes, and oomycetes. Proc. Natl. Acad. Sci. U.S.A. 84: 5823-5827.
Hall, J. D. and Delwiche, C. F. 2007. In the shadow of giants: systematics of the charophyte green algae. In: Unravelling the algae: the past, present, and future of algal systematics. pp. 155-169. Brodie, J. & Lewis, J. Eds., Taylor and Francis.
Hall, J. D., Karol, K. G., McCourt, R. M. and Delwiche, C. F. 2008a. Phylogeny of the conjugating green algae based on chloroplast and mitochondrial nucleotide sequence data. J. Phycol. 44: 467-477.
Hall, J. D., McCourt, R. M. and Delwiche, C. F. 2008b. Patterns of cell division in the filamentous Desmidiaceae, close green algal relatives of land plants. Am. J. Bot. 95: 643-654.
Hallick, R. B., Hong, L., Drager, R. G., Favreau, M. R., Monfort, A., Orsat, B., Spielmann, A. and Stutz, E. 1993. Complete sequence of Euglena gracilis chloroplast DNA. Nucleic Acids Res. 21: 3537-3544.
Hallmann, A. 2003. Extracellular matrix and sex-inducing pheromone in Volvox. Int. Rev. Cytol. 227: 131-182.
Hämmerling, J. 1953. Nucleo-cytoplasmic relationships in the development of Acetabularia. Int. Rev. Cytol. 2: 475-498.
Hanagata, N. and Chihara, M. 1999. Coenocystis inconstans, a new species of bark-inhabiting green algae (Chlorophyceae, Chlorophyta). J. Jpn. Bot. 74: 204-211.
Händeler, K., Grzymbowski, Y. P., Krug, P. J. and Wägele, H. 2009. Functional chloroplasts in metazoan cells – a unique evolutionary strategy in animal life. Front. Zool. 6: 28.
Händeler, K., Wägele, H., Wahrmund, U., Rüdinger, M. and Knoop, V. 2010. Slugs’ last meals: molecular identification of sequestered chloroplasts from different algal origins in Sacoglossa (Opisthobranchia, Gastropoda). Mol. Ecol. Resour. 10: 968–978.
Hannaert, V., Saavedra, E., Duffieux, F., Szikora, J.-P., Rigden, D. J., Michels, P. A. M. and Opperdoes, F. R. 2003. Plant-like traits associated with metabolism of Trypanosoma parasites. Proc. Natl. Acad. Sci. U.S.A. 100: 1067-1071.
Hanyuda, T., Wakana, I., Arai, S., Miyaji, K., Watano, Y. and Ueda, K. 2002. Phylogenetic relationships within Cladophorales (Ulvophyceae, Chlorophyta) inferred from 18S rRNA gene sequences, with special reference to Aegagropila linnaei. J. Phycol. 38: 564-571.
Harris, E. H. 2001. Chlamydomonas as a model organism. Annu. Rev. Plant Physiol. Plant Mol. Biol. 52: 363-406.
Hasegawa, T., Miyashita, H., Kawachi, M., Ikemoto, H., Kurano, N., Miyachi, S. and Chihara, M. 1996. Prasinoderma coloniale gen. et sp. nov., a new pelagic coccoid prasinophyte from the western Pacific ocean. Phycologia 35: 170-176.
Hayden, H. S. and Waaland, J. R. 2002. Phylogenetic systematics of the Ulvaceae (Ulvales, Ulvophyceae) using chloroplast and nuclear DNA sequences. J. Phycol. 38: 1200-1212.
Hayden, H. S. and Waaland, J. R. 2004. A molecular systematic study of Ulva (Ulvaceae, Ulvales) from the northeast Pacific. Phycologia 43: 364-382.
Heckman, D. S., Geiser, D. M., Eidell, B. R., Stauffer, R. L., Kardos, N. L. and Hedges, S. B. 2001. Molecular evidence for the early colonization of land by fungi and plants. Science 293: 1129-1133.
Hedges, S. B., Blair, J. E., Venturi, M. L. and Shoe, J. L. 2004. A molecular timescale of eukaryote evolution and the rise of complex multicellular life. BMC Evol. Biol. 4: 2.
Heesch, S., Broom, J. E. S., Neill, K. F., Farr, T. J., Dalen, J. L. and Nelson, W. A. 2009. Ulva, Umbraulva and Gemina: genetic survey of New Zealand taxa reveals diversity and introduced species. Eur. J. Phycol. 44: 143-154.
Henley, W. J., Hironaka, J. L., Guillou, L., Buchheim, M. A., Buchheim, J. A., Fawley, M. W. and Fawley, K. P. 2004. Phylogenetic analysis of the ‘Nannochloris-like’ algae and diagnoses of Picochlorum oklahomensis gen. et sp. nov. (Trebouxiophyceae, Chlorophyta). Phycologia 43: 641-652.
Henschel, K., Kofuji, R., Hasebe, M., Saedler, H., Munster, T. and Theissen, G. 2002. Two ancient classes of MIKC-type MADS-box genes are present in the moss Physcomitrella patens. Mol. Biol. Evol. 19: 801-814.
Hepperle, D., Hegewald, E. and Krienitz, L. 2000. Phylogenetic position of the Oocystaceae (Chlorophyta). J. Phycol. 36: 590-595.
Herron, M. D., Hackett, J. D., Aylward, F. O. and Michod, R. E. 2009. Triassic origin and early radiation of multicellular volvocine algae. Proc. Natl. Acad. Sci. U.S.A. 106: 3254-3258.
Herron, M. D. and Michod, R. E. 2008. Evolution of complexity in the volvocine algae: Transitions in individuality through Darwin’s eye. Evolution 62: 436-451.
Hillis-Colinvaux, L. 1984. Systematics of the Siphonales. In: Systematics of the green algae. pp. 271-291. Irvine, D. E. G. & John, D. M. Eds., Academic Press, London.
Hoops, H. J. 1993. Flagellar, cellular and organismal polarity in Volvox carteri. J. Cell Sci. 104: 105-117.
Hori, H., Lim, B.-L. and Osawa, S. 1985. Evolution of green plants as deduced from 5S rRNA sequences. Proc. Natl. Acad. Sci. U.S.A. 82: 820-823.
Hori, H. and Osawa, S. 1987. Origin and evolution of organisms as deduced from 5S ribosomal RNA sequences. Mol. Biol. Evol. 4: 445-472.
Hoshina, R. and Imamura, N. 2008. Multiple origins of the symbioses in Paramecium bursaria. Protist 159: 53-63.
Hua, J., Smith, D. R., Borza, T. and Lee, R. W. 2011. Similar relative mutation rates in the three genetic compartments of Mesostigma and Chlamydomonas. Protist: In press.
Huang, J. L., Mullapudi, N., Lancto, C. A., Scott, M., Abrahamsen, M. S. and Kissinger, J. C. 2004. Phylogenomic evidence supports past endosymbiosis, intracellular and horizontal gene transfer in Cryptosporidium parvum. Genome Biology 5: R88.
Huss, V. A. R., Ciniglia, C., Cennamo, P., Cozzolino, S., Pinto, G. and Pollio, A. 2002. Phylogenetic relationships and taxonomic position of Chlorella-like isolates from low pH environments (pH < 3.0). BMC Evol. Biol. 2: 13.
Huss, V. A. R., Frank, C., Hartmann, E. C., Hirmer, M., Kloboucek, A., Seidel, B. M., Wenzeler, P. and Kessler, E. 1999. Biochemical taxonomy and molecular phylogeny of the genus Chlorella sensu lato (Chlorophyta). J. Phycol. 35: 587-598.
Huss, V. A. R. and Sogin, M. L. 1990. Phylogenetic position of some Chlorella species within the Chlorococcales based upon complete small-subunit ribosomal RNA sequences. J. Mol. Evol. 31: 432-442.
Ichihara, K., Arai, S., Uchimura, M., Fay, E. J., Ebata, H., Hiraoka, M. and Shimada, S. 2009. New species of freshwater Ulva, Ulva limnetica (Ulvales, Ulvophyceae) from the Ryukyu Islands, Japan. Phycol. Res. 57: 94-103.
Ishida, K., Yabuki, A. and Ota, S. 2007. The chlorarachniophytes: evolution and classification. In: Unravelling the algae: the past, present, and future of algal systematics. pp. 171-182. Brodie, J. & Lewis, J. Eds., CRC Press, Taylor and Francis, Boca Raton.
Iyengar, M. O. P. and Desikachary, T. V. 1981. Volvocales. Indian Council of Agricultural Research, New Delhi.
Jacobshagen, S. and Schnarrenberger, C. 1990. Two class I aldolases in Klebsormidium flaccidum (Charophyceae): an evolutionary link from chlorophytes to higher plants. J. Phycol. 26: 312-317.
Janouškovec, J., Horák, A., Oborník, M., Lukeš, J. and Keeling, P. J. 2010. A common red algal origin of the apicomplexan, dinoflagellate, and heterokont plastids. Proc. Natl. Acad. Sci. U.S.A. 107: 10949-10954.
Jeffrey, C. 1971. Thallophytes and kingdoms – a critique. Kew Bull. Addit. Ser. 25: 291-299.
Jeffrey, C. 1982. Kingdoms, codes and classification. Kew Bull. 37: 403-416.
Jiao, H. S., Hicks, A., Simpson, C. and Stern, D. B. 2004. Short dispersed repeats in the Chlamydomonas chloroplast genome are collocated with sites for mRNA 3′ end formation. Curr. Genet. 45: 311-322.
Jobson, R. W. and Qiu, Y. L. 2008. Did RNA editing in plant organellar genomes originate under natural selection or through genetic drift? Biol. Direct 3: 43.
John, D. M., Whitton, B. A. and Brook, A. J. 2002. The Freshwater Algal Flora of the British Isles. An Identification Guide to Freshwater and Terrestrial Algae. Cambridge University Press, Cambridge.
Johnson, J. L., Fawley, M. W. and Fawley, K. P. 2007. The diversity of Scenedesmus and Desmodesmus (Chlorophyceae) in Itasca State Park, Minnesota, USA. Phycologia 46: 214-229.
Johnson, P. W. and Sieburth, J. M. 1982. In-situ morphology and occurrence of eucaryotic phototrophs of bacterial size in the picoplankton of estuarine and oceanic waters. Blackwell Publishing Ltd, pp. 318-327.
Joubert, J. J. and Rijkenberg, F. H. J. 1971. Parasitic green algae. Annu. Rev. Phytopathol. 9: 45-64.
Jouenne, F., Eikrem, W., Le Gall, F., Marie, D., Johnsen, G. and Vaulot, D. 2011. Prasinoderma singularis sp. nov. (Prasinophyceae, Chlorophyta), a solitary coccoid prasinophyte from the South-East Pacific Ocean. Protist 162: 70-84.
Jousson, O., Pawlowski, J., Zaninetti, L., Zechman, F. W., Dini, F., Di Guiseppe, G., Woodfield, R., Millar, A. and Meinesz, A. 2000. Invasive alga reaches California: The alga has been identified that threatens to smother Californian coastal ecosystems. Nature 408: 157-158.
Kamikawa, R., Inagaki, Y. and Sako, Y. 2008. Direct phylogenetic evidence for lateral transfer of elongation factor-like gene. Proc. Natl. Acad. Sci. U.S.A. 105: 6965-6969.
Kantz, T. S., Theriot, E. C., Zimmer, E. A. and Chapman, R. L. 1990. The Pleurastrophyceae and Micromonadophyceae: a cladistics analysis of nuclear rRNA sequence data. J. Phycol. 26: 711-721.
Karakashian, S. J. and Karakashian, M. W. 1965. Evolution and symbiosis in the genus Chlorella and related algae. Evolution 19: 368-377.
Karol, K. G., McCourt, R. M., Cimino, M. T. and Delwiche, C. F. 2001. The closest living relatives of land plants. Science 294: 2351-2353.
Karsten, U., Friedl, T., Schumann, R., Hoyer, K. and Lembcke, S. 2005. Mycosporine-like amino acids and phylogenies in green algae: Prasiola and its relatives from the Trebouxiophyceae (Chlorophyta). J. Phycol. 41: 557-566.
Katana, A., Kwiatowski, J., Spalik, K., Zakrys, B., Szalacha, E. and Szymanska, H. 2001. Phylogenetic position of Koliella (Chlorophyta) as inferred from nuclear and chloroplast small subunit rDNA. J. Phycol. 37: 443-451.
Keeling, P. J. 2004. Diversity and evolutionary history of plastids and their hosts. Am. J. Bot. 91: 1481-1493.
Keeling, P. J. 2010. The endosymbiotic origin, diversification and fate of plastids. Philos. Trans. R. Soc. Lond. B Biol. Sci. 365: 729-748.
Keeling, P. J. and Inagaki, Y. 2004. A class of eukaryotic GTPase with a punctate distribution suggesting multiple functional replacements of translation elongation factor 1α. Proc. Natl. Acad. Sci. U.S.A. 101: 15380-15385.
Keeling, P. J. and Palmer, J. D. 2008. Horizontal gene transfer in eukaryotic evolution. Nat. Rev. Genet. 9: 605-618.
Keller, A., Schleicher, T., Forster, F., Ruderisch, B., Dandekar, T., Muller, T. and Wolf, M. 2008. ITS2 data corroborate a monophyletic chlorophycean DO-group (Sphaeropleales). BMC Evol. Biol. 8: 218.
Kenrick, P. and Crane, P. R. 1997. The origin and early evolution of plants on land. Nature 389: 33-39.
Kerney, R., Kim, E., Hangarter, R. P., Heiss, A. A., Bishop, C. D. and Hall, B. K. 2011. Intracellular invasion of green algae in a salamander host. Proc. Natl. Acad. Sci. U.S.A.: (in press).
Kim, G. H., Klotchkova, T. A. and Kang, Y. M. 2001. Life without a cell membrane: regeneration of protoplasts from disintegrated cells of the marine green alga Bryopsis plumosa. J. Cell Sci. 114: 2009-2014.
Kim, J. H., Kim, Y. H., Cho, G. Y. and Boo, S. M. 2006. Plastid rbcL gene phylogeny of the genus Spirogyra (chlorophyta, zygnemataceae) from Korea. Korean J. Genet. 28: 295-303.
Kirk, D. L. 1998. Volvox: Molecular-genetic origins of multicellularity and cellular differentiation. Cambridge University Press, Cambridge.
Kirk, D. L. 2003. Seeking the ultimate and proximate causes of Volvox multicellularity and cellular differentiation. Integr. Comp. Biol. 43: 247-253.
Kirk, D. L. 2005. A twelve-step program for evolving multicellularity and a division of labor. BioEssays 27: 299-310.
Kirk, D. L., Birchem, R. and King, N. 1986. The extracellular matrix of Volvox: a comparative study and proposed system of nomenclature. J. Cell Sci. 80: 207-231.
Kirk, D. L. and Kirk, M. M. 1986. Heat shock elicits production of sexual inducer in Volvox. Science 231: 51-54.
Kirk, M. M. and Kirk, D. L. 1985. Translational regulation of protein synthesis, in response to light, at a critical stage of Volvox development. Cell 41: 419-428.
Kirk, M. M., Ransick, A., McRae, S. E. and Kirk, D. L. 1993. The relationship between cell size and cell fate in Volvox carteri. J. Cell Biol. 123: 191-208.
Kirk, M. M., Stark, K., Miller, S. M., Muller, W., Taillon, B. E., Gruber, H., Schmitt, R. and Kirk, D. L. 1999. regA, a Volvox gene that plays a central role in germ-soma differentiation, encodes a novel regulatory protein. Development 126: 639-647.
Knoll, A. H. 2003. Life on a Young Planet: The First Three Billion Years of Evolution on Earth. Princeton University Press, Princeton, NJ.
Koonin, E. V. 2005. Orthologs, paralogs, and evolutionary genomics. Annu. Rev. Genet. 39: 309-338.
Kovacevic, G., Franjevic, D., Jelencic, B. and Kalafatic, M. 2010. Isolation and cultivation of endosymbiotic algae from green Hydra and phylogenetic analysis of 18S rDNA sequences. Folia Biol. (Kraków) 58: 135-143.
Kraft, L. G. K., Kraft, G. T. and Waller, R. F. 2010. Investigations into southern australian Ulva (Ulvophyceae, Chlorophyta) taxonomy and molecular phylogeny indicate both cosmopolitanism and endemic cryptic species. J. Phycol. 46: 1257-1277.
Kranz, H. D., Miks, D., Siegler, M. L., Capesius, I., Sensen, C. W. and Huss, V. A. R. 1995. The origin of land plants: phylogenetic relationships among charophytes, bryophytes, and vascular plants inferred from complete small-subunit ribosomal RNA gene sequences. J. Mol. Evol. 41: 74-84.
Kreimer, G. 2009. The green algal eyespot apparatus: a primordial visual system and more? Curr. Genet. 55: 19-43.
Krienitz, L., Bock, C., Dadheech, P. K. and Pröschold, T. 2011. Taxonomic reassessment of the genus Mychonastes (Chlorophyceae, Chlorophyta) including the description of eight new species. Phycologia 50: 89-106.
Krienitz, L., Bock, C., Luo, W. and Pröschold, T. 2010. Polyphyletic origin of the Dictyosphaerium morphotype within Chlorellaceae (Trebouxiophyceae). J. Phycol. 46: 559-563.
Krienitz, L., Hegewald, E., Hepperle, D. and Wolf, M. 2003. The systematics of coccoid green algae: 18S rRNA gene sequence data versus morphology. Biologia 58: 437-446.
Krienitz, L., Ustinova, I., Friedl, T. and Huss, V. A. R. 2001. Traditional generic concepts versus 18S rRNA gene phylogeny in the green algal family Selenastraceae (Chlorophyceae, Chlorophyta). J. Phycol. 37: 852-865.
La Claire, J. W. 1992. Contractile movements in the algae: the Siphonocladales as model systems. In: The Cytoskeleton of the Algae. pp. 239-253. Menzel, D. Ed., CRC Press, Boca Raton.
Laflamme, M. and Lee, R. W. 2003. Mitochondrial genome conformation among CW-group chlorophycean algae. J. Phycol. 39: 213-220.
Lam, D. W. and Zechman, F. W. 2006. Phylogenetic analyses of the Bryopsidales (Ulvophyceae, Chlorophyta) based on Rubisco large subunit gene sequences. J. Phycol. 42: 669-678.
Lang, B. F., Gray, M. W. and Burger, G. 1999. Mitochondrial genome evolution and the origin of eukaryotes. Annu. Rev. Genet. 33: 351-397.
Lang, D., Zimmer, A. D., Rensing, S. A. and Reski, R. 2008. Exploring plant biodiversity: the Physcomitrella genome and beyond. Trends Plant Sci. 13: 542-549.
Lapointe, B. E., Barile, P. J., Littler, M. M., Littler, D. S., Bedford, B. J. and Gasque, C. 2005. Macroalgal blooms on southeast Florida coral reefs I. Nutrient stoichiometry of the invasive green alga Codium isthmocladum in the wider Caribbean indicates nutrient enrichment. Harmful Algae 4: 1092-1105.
Latasa, M., Scharek, R., Le Gall, F. and Guillou, L. 2004. Pigment suites and taxonomic groups in Prasinophyceae. J. Phycol. 40: 1149-1155.
* Laurin-Lemay, S., Brinkmann, H. and Philippe, H. 2012. Origin of land plants revisited in the light of sequence contamination and missing data. Curr. Biol. 22: R593-R594.
Leliaert, F., De Clerck, O., Verbruggen, H., Boedeker, C. and Coppejans, E. 2007. Molecular phylogeny of the Siphonocladales (Chlorophyta: Cladophorophyceae). Mol. Phylogenet. Evol. 44: 1237-1256.
Leliaert, F., Rousseau, F., De Reviers, B. and Coppejans, E. 2003. Phylogeny of the Cladophorophyceae (Chlorophyta) inferred from partial LSU rRNA gene sequences: is the recognition of a separate order Siphonocladales justified? Eur. J. Phycol. 38: 233-246.
Leliaert, F., Rueness, J., Boedeker, C., Maggs, C. A., Cocquyt, E., Verbruggen, H. and De Clerck, O. 2009a. Systematics of the marine microfilamentous green algae Uronema curvatum and Urospora microscopica (Chlorophyta). Eur. J. Phycol. 44: 487-496.
Leliaert, F., Verbruggen, H., Wysor, B. and De Clerck, O. 2009b. DNA taxonomy in morphologically plastic taxa: Algorithmic species delimitation in the Boodlea complex (Chlorophyta: Cladophorales). Mol. Phylogenet. Evol. 53: 122-133.
Leliaert, F., Zhang, X. W., Ye, N. H., Malta, E., Engelen, A. H., Mineur, F., Verbruggen, H. and De Clerck, O. 2009c. Identity of the Qingdao algal bloom. Phycol. Res. 57: 147-151.
Lemieux, C., Otis, C. and Turmel, M. 2000. Ancestral chloroplast genome in Mesostigma viride reveals an early branch of green plant evolution. Nature 403: 649-652.
Lemieux, C., Otis, C. and Turmel, M. 2007. A clade uniting the green algae Mesostigma viride and Chlorokybus atmophyticus represents the deepest branch of the Streptophyta in chloroplast genome-based phylogenies. BMC Biology 5: 2.
Lenz, H., Rudinger, M., Volkmar, U., Fischer, S., Herres, S., Grewe, F. and Knoop, V. 2010. Introducing the plant RNA editing prediction and analysis computer tool PREPACT and an update on RNA editing site nomenclature. Curr. Genet. 56: 189-201.
Lepère, C., Vaulot, D. and Scanlan, D. J. 2009. Photosynthetic picoeukaryote community structure in the South East Pacific Ocean encompassing the most oligotrophic waters on Earth. Environ. Microbiol. 11: 3105-3117.
Letsch, M. R., Muller-Parker, G., Friedl, T. and Lewis, L. A. 2009. Elliptochloris marina sp. nov. (Trebouxiophyceae, Chlorophyta), symbiotic green alga of the temperate pacific sea anemones Anthopleura xanthogrammica and A. elegantissima (Anthozoa, Cnidaria). J. Phycol. 45: 1127-1135.
Lewis, L. A. 1997. Diversity and phylogenetic placement of Bracteacoccus Tereg (Chlorophyceae, Chlorophyta) based on 18S ribosomal RNA gene sequence data. J. Phycol. 33: 279-285.
Lewis, L. A. and Lewis, P. O. 2005. Unearthing the molecular phylodiversity of desert soil green algae (Chlorophyta). Syst. Biol. 54: 936-947.
Lewis, L. A. and McCourt, R. M. 2004. Green algae and the origin of land plants. Am. J. Bot. 91: 1535-1556.
Lewis, L. A. and Muller-Parker, G. 2004. Phylogenetic placement of “Zoochlorellae” (Chlorophyta), algal symbiont of the temperate sea anemone Anthopleura elegantissima. Biol. Bull. 207: 87-92.
Lewis, L. A., Wilcox, L. W., Fuerst, P. A. and Floyd, G. L. 1992. Concordance of molecular and ultrastructural data in the study of zoosporic chlorococcalean green algae. J. Phycol. 28: 375-380.
Li, W. 1994. Primary production of prochlorophytes, cyanobacteria, and eucaryotic ultraphytoplankton: Measurements from flow cytometric sorting. Limnol. Oceanogr. 39: 169-175.
Li, W. Y., Liu, B., Yu, L. J., Feng, D. R., Wang, H. B. and Wang, J. F. 2009. Phylogenetic analysis, structural evolution and functional divergence of the 12-oxo-phytodienoate acid reductase gene family in plants. BMC Evol. Biol. 9: 90.
Ligrone, R., Carafa, A., Duckett, J., Renzaglia, K. and Ruel, K. 2008. Immunocytochemical detection of lignin-related epitopes in cell walls in bryophytes and the charalean alga Nitella. Plant Syst. Evol. 270: 257-272.
Lindstrom, S. C. and Hanic, L. A. 2005. The phylogeny of North American Urospora (Ulotrichales, Chlorophyta) based on sequence analysis of nuclear ribosomal genes, introns and spacers. Phycologia 44: 194-201.
Lindstrom, S. C., Hanic, L. A. and Golden, L. 2006. Studies of the green alga Percursaria dawsonii (=Blidingia dawsonii comb. nov., Kornmanniaceae, Ulvales) in British Columbia. Phycol. Res. 54: 40-56.
Lokhorst, G. M. and Rongen, G. P. J. 1994. Comparative ultrastructural studies of division processes in terrestrial green alga Leptosira erumpens (Deason & Bold) Lukesova confirm the ordinal status of the Pleurastrales. Crypt. Bot. 4: 394-409.
Lokhorst, G. M., Sluiman, H. J. and Star, W. 1988. The ultrastructure of mitosis and cytokinesis in the sarcinoid Chlorokybus atmophyticus (Chlorophyta, Charophyceae) revealed by rapid freeze fixation and freeze substitution. J. Phycol. 24: 237-248.
Lokhorst, G. M. and Star, W. 1999. The flagellar apparatus structure in Microspora (Chlorophyceae) confirms a close evolutionary relationship with unicellular green algae. Plant Syst. Evol. 217: 11-30.
López-Bautista, J. M. and Chapman, R. L. 2003. Phylogenetic affinities of the Trentepohliales inferred from small-subunit rDNA. Int. J. Syst. Evol. Microbiol. 53: 2099-2106.
López-Bautista, J. M., Rindi, F. and Guiry, M. D. 2006. Molecular systematics of the subaerial green algal order Trentepohliales: an assessment based on morphological and molecular data. Int. J. Syst. Evol. Microbiol. 56: 1709-1715.
Lopez-Garcia, P., Rodriguez-Valera, F., Pedros-Alio, C. and Moreira, D. 2001. Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature 409: 603-607.
Loughnane, C. J., McIvor, L. M., Rindi, F., Stengel, D. B. and Guiry, M. D. 2008. Morphology, rbcL phylogeny and distribution of distromatic Ulva (Ulvophyceae, Chlorophyta) in Ireland and southern Britain. Phycologia 47: 416-429.
Lü, F., Xü, W., Tian, C., Wang, G., Niu, J., Pan, G. and Hu, S. 2011. The Bryopsis hypnoides plastid genome: multimeric forms and complete nucleotide sequence. PLoS One 6: e14663.
Luo, W., Proschold, T., Bock, C. and Krienitz, L. 2010. Generic concept in Chlorella-related coccoid green algae (Chlorophyta, Trebouxiophyceae). Plant Biology 12: 545-553.
Lynch, M. and Conery, J. S. 2003. The origins of genome complexity. Science 302: 1401-1404.
Lynch, M., Koskella, B. and Schaack, S. 2006. Mutation pressure and the evolution of organelle genomic architecture. Science 311: 1727-1730.
Malek, O., Lattig, K., Hiesel, R., Brennicke, A. and Knoop, V. 1996. RNA editing in bryophytes and a molecular phylogeny of land plants. EMBO J. 15: 1403-1411.
Malkin, S. Y., Dove, A., Depew, D., Smith, R. E., Guildford, S. J. and Hecky, R. E. 2010. Spatiotemporal patterns of water quality in Lake Ontario and their implications for nuisance growth of Cladophora. J. Gt. Lakes Res. 36: 477-489.
Mallet, M. A. and Lee, R. W. 2006. Identification of three distinct Polytomella lineages based on mitochondrial DNA features. J. Eukaryot. Microbiol. 53: 79-84.
Mandoli, D. F. 1998. Elaboration of body plan and phase change during development of Acetabularia: How is the complex architecture of a giant unicell built? Annu. Rev. Plant Physiol. Plant Mol. Biol. 49: 173-198.
Manhart, J. R. and Palmer, J. D. 1990. The gain of two chloroplast tRNA introns marks the green algal ancestors of land plants. Nature 345: 268-270.
Manton, I. 1964. Observations on fine structure of zoospore and young germling of Stigeoclonium. J. Exp. Bot. 15: 399-&.
Manton, I. 1967. Electron microscopical observations on a clone of Monomastix Scherffel in culture. Nova Hedwigia 14: 1-11.
Marchant, H. J. 1977. Colony formation and inversion in the green alga Eudorina elegans. Protoplasma 93: 325-339.
Marchant, H. J. and Pickett-Heaps, J. D. 1973. Mitosis and cytokinesis in Coleochaete scutata. J. Phycol. 9: 461-471.
Marie, D., Shi, X. L., Rigaut-Jalabert, F. and Vaulot, D. 2010. Use of flow cytometric sorting to better assess the diversity of small photosynthetic eukaryotes in the English Channel. FEMS Microbiol. Ecol. 72: 165-178.
Marie, D., Zhu, F., Balague, V., Ras, J. and Vaulot, D. 2006. Eukaryotic picoplankton communities of the Mediterranean Sea in summer assessed by molecular approaches (DGGE, TTGE, QPCR). FEMS Microbiol. Ecol. 55: 403-415.
Marin, B. and Melkonian, M. 1994. Flagellar hairs in prasinophytes (Chlorophyta): Ultrastructure and distribution on the flagellar surface. J. Phycol. 30: 659-678.
Marin, B. and Melkonian, M. 1999. Mesostigmatophyceae, a new class of streptophyte green algae revealed by SSU rRNA sequence comparisons. Protist 150: 399-417.
Marin, B. and Melkonian, M. 2010. Molecular phylogeny and classification of the Mamiellophyceae class. nov. (Chlorophyta) based on sequence comparisons of the nuclear- and plastid-encoded rRNA operons. Protist 161: 304-336.
* Marin, B. 2012. Nested in the Chlorellales or independent class? Phylogeny and classification of the Pedinophyceae (Viridiplantae) revealed by molecular phylogenetic analyses of complete nuclear and plastid-encoded rRNA operons. Protist 163: 778–805
Massana, R., Balague, V., Guillou, L. and Pedros-Alio, C. 2004. Picoeukaryotic diversity in an oligotrophic coastal site studied by molecular and culturing approaches. FEMS Microbiol. Ecol. 50: 231-243.
Massjuk, N. P. 2006. Chlorodendrophyceae class. nov. (Chlorophyta, Viridiplantae) in the Ukrainian flora: I. The volume, phylogenetic relations and taxonomical status. Ukr. Bot. J. 63: 601-614 (in Ukrainian).
Matsumoto, T., Shinozaki, F., Chikuni, T., Yabuki, A., Takishita, K., Kawachi, M., Nakayama, T., Inouye, I., Hashimoto, T. and Inagaki, Y. 2011. Green-colored plastids in the dinoflagellate genus Lepidodinium are of core chlorophyte origin. Protist 162 268-276.
Matsuzaki, R., Nakada, T., Hara, Y. and Nozaki, H. 2010. Light and electron microscopy and molecular phylogenetic analyses of Chloromonas pseudoplatyrhyncha (Volvocales, Chlorophyceae). Phycol. Res. 58: 202-209.
Mattox, K. R. and Stewart, K. D. 1984. Classification of the green algae: a concept based on comparative cytology. In: Systematics of the green algae. pp. 29-72. Irvine, D. E. G. & John, D. M. Eds., Academic Press, London.
Maul, J. E., Lilly, J. W., Cui, L. Y., dePamphilis, C. W., Miller, W., Harris, E. H. and Stern, D. B. 2002. The Chlamydomonas reinhardtti plastid chromosome: Islands of genes in a sea of repeats. Plant Cell 14: 2659-2679.
McCourt, R. M. 1995. Green algal phylogeny. Trends Ecol. Evol. 10: 159-163.
McCourt, R. M., Delwiche, C. F. and Karol, K. G. 2004. Charophyte algae and land plant origins. Trends Ecol. Evol. 19: 661-666.
McCourt, R. M., Karol, K. G., Bell, J., Helm-Bychowski, K. M., Grajewska, A., Wojciechowski, M. F. and Hoshaw, R. W. 2000. Phylogeny of the conjugating green algae (Zygnemophyceae) based on rbcL sequences. J. Phycol. 36: 747-758.
McCourt, R. M., Karol, K. G., Casanova, M. T. and Feist, M. 1999. Monophyly of genera and species of Characeae based on rbcL sequences, with special reference to Australian and European Lychnothamnus barbatus (Characeae: Charophyceae). Aust. J. Bot. 47: 361-369.
McCourt, R. M., Karol, K. G., Guerlesquin, M. and Feist, M. 1996. Phylogeny of extant genera in the family Characeae (Charales, Charophyceae) based on rbcL sequences and morphology. Am. J. Bot. 83: 125-131.
McDonald, S. M., Plant, J. N. and Worden, A. Z. 2010. The mixed lineage nature of nitrogen transport and assimilation in marine eukaryotic phytoplankton: A case study of Micromonas. Mol. Biol. Evol. 27: 2268-2283.
McManus, H. A. and Lewis, L. A. 2005. Molecular phylogenetics, morphological variation and colony-form evolution in the family Hydrodictyaceae (Sphaeropleales, Chlorophyta). Phycologia 44: 582-595.
McManus, H. A. and Lewis, L. A. 2011. Molecular phylogenetic relationships in the freshwater family Hydrodictyaceae (Sphaeropleales, Chlorophyceae), with an emphasis on Pediastrum duplex. J. Phycol. 47: 152-163.
McNaughton, E. E. and Goff, L. J. 1990. The role of microtubules in establishing nuclear spatial patterns in multinucleate green algae. Protoplasma 157: 19-37.
Mei, H., Luo, W., Liu, G. X. and Hu, Z. Y. 2007. Phylogeny of Oedogoniales (Chlorophyceae, Chlorophyta) inferred from 18S rDNA sequences with emphasis on the relationships in the genus Oedogonium based on ITS-2 sequences. Plant Syst. Evol. 265: 179-191.
Meinesz, A. and Hesse, B. 1991. Introduction of the tropical alga Caulerpa taxifolia and its invasion of the northwestern Mediterranean. Oceanol. Acta 14: 415-426.
Meissner, M., Stark, K., Cresnar, B., Kirk, D. L. and Schmitt, R. 1999. Volvox germline-specific genes that are putative targets of RegA repression encode chloroplast proteins. Curr. Genet. 36: 363-370.
Melkonian, M. 1975. Fine structure of zoospores of Fritschiella tuberosa Iyeng (Chaetophorineae, Chlorophyceae) with special reference to flagellar apparatus. Protoplasma 86: 391-404.
Melkonian, M. 1982. Structural and evolutionary aspects of the flagellar apparatus in green algae and land plants. Taxon 31: 255-265.
Melkonian, M. 1984. Flagellar apparatus ultrastructure in relation to green algal classification. In: Systematics of the green algae. pp. 73–120. Irvine, D. E. G. & John, D. M. Eds., Academic Press, London.
Melkonian, M. 1989. Flagellar apparatus ultrastructure in Mesostigma viride (Prasinophyceae). Plant Syst. Evol. 164: 93-122.
Melkonian, M. 1990a. Chlorophyte orders of uncertain affinities: order Pedinomonadales. In: Handbook of Protoctista. The structure, cultivation, habitats and life histories of the eukaryotic microorganisms and their descendants exclusive of animals, plants and fungi. pp. 649–651. Margulis, L., Corliss, J. O., Melkonian, M. & Chapman, D. J. Eds., Jones and Bartlett Publishers, Boston.
Melkonian, M. 1990b. Phylum Chlorophyta. Class Prasinophyceae. In: Handbook of Protoctista. The structure, cultivation, habitats and life histories of the eukaryotic microorganisms and their descendants exclusive of animals, plants and fungi. pp. 600-607. Margulis, L., Corliss, J. O., Melkonian, M. & Chapman, D. J. Eds., Jones and Bartlett Publishers, Boston.
Menzel, D. 1987. The cytoskeleton of the giant coenocytic green-alga Caulerpa visualized by immunocytochemistry. Protoplasma 139: 71-76.
Menzel, D. 1988. How do giant plant cells cope with injury? – The wound response in siphonous green algae. Protoplasma 144: 73-91.
Menzel, D. 1994. Cell differentiation and the cytoskeleton in Acetabularia. New Phytol. 128: 369-393.
Merchant, S. S., Prochnik, S. E., Vallon, O., Harris, E. H., Karpowicz, S. J., et al. 2007. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318: 245-251.
Metzker, M. L. 2010. Sequencing technologies: the next generation. Nat. Rev. Genet. 11: 31-46.
Michaelis, G., Vahrenholz, C. and Pratje, E. 1990. Mitochondrial DNA of Chlamydomonas reinhardtii: the gene for apocytochrome b and the complete functional map of the 15.8 kb DNA. Mol. Gen. Genet. 223: 211-216.
Mikhailyuk, T. I., Sluiman, H. J., Massalski, A., Mudimu, O., Demchenko, E. M., Kondratyuk, S. Y. and Friedl, T. 2008. New streptophyte green algae from terrestrial habitats and an assessment of the genus Interfilum (Klebsormidiophyceae, Streptophyta). J. Phycol. 44: 1586-1603.
Mine, I., Menzel, D. and Okuda, K. 2008. Morphogenesis in giant-celled algae. Int. Rev. Cell Mol. Biol. 266: 37-83.
Mine, I., Okuda, K. and Menzel, D. 2001. Poly(A)+ RNA during vegetative development of Acetabularia peniculus. Protoplasma 216: 56-65.
Mishler, B. D., Lewis, L. A., Buchheim, M. A., Renzaglia, K. S., Garbary, D. J., Delwiche, C. F., Zechman, F. W., Kantz, T. S. and Chapman, R. L. 1992. Phylogenetic relationships of the “green algae” and “bryophytes”. Ann. Mo. Bot. Gard. 81: 451-483.
Moestrup, Ø. 1978. On the phylogenetic validity of the flagellar apparatus in green algae and other chlorophyll a and b containing plants. Biosystems 10: 117-144.
Moestrup, Ø. 1991. Further studies of presumedly primitive green algae, including the description of Pedinophyceae class. nov. and Resultor gen. nov. J. Phycol. 27: 119-133.
Moestrup, Ø., Inouye, I. and Hori, T. 2003. Ultrastructural studies on Cymbomonas tetramitiformis (Prasinophyceae). I. General structure, scale microstructure, and ontogeny. Can. J. Bot. 81: 657-671.
Moon-van der Staay, S. Y., De Wachter, R. and Vaulot, D. 2001. Oceanic 18S rDNA sequences from picoplankton reveal unsuspected eukaryotic diversity. Nature 409: 607-610.
Moore, C. E. and Archibald, J. M. 2009. Nucleomorph genomes. Annu. Rev. Genet. 43: 251-264.
Motomura, T. 1996. Cell cycle analysis in a multinucleate green alga, Boergesenia forbesii (Siphonocladales, Chlorophyta). Phycol. Res. 44 11-17.
Moustafa, A., Beszteri, B., Maier, U. G., Bowler, C., Valentin, K. and Bhattacharya, D. 2009. Genomic footprints of a cryptic plastid endosymbiosis in diatoms. Science 324: 1724-1726.
Mujer, C. V., Andrews, D. L., Manhart, J. R., Pierce, S. K. and Rumpho, M. E. 1996. Chloroplast genes are expressed during intracellular symbiotic association of Vaucheria litorea plastids with the sea slug Elysia chlorotica. Proc. Natl. Acad. Sci. U.S.A. 93: 12333-12338.
Mukherjee, K., Brocchieri, L. and Burglin, T. R. 2009. A comprehensive classification and evolutionary analysis of plant homeobox genes. Mol. Biol. Evol. 26: 2775-2794.
Müller, T., Rahmann, S., Dandekar, T. and Wolf, M. 2004. Accurate and robust phylogeny estimation based on profile distances: a study of the Chlorophyceae (Chlorophyta). BMC Evol. Biol. 4: 20.
Muramoto, K., Nakada, T., Shitara, T., Hara, Y. and Nozaki, H. 2010. Re-examination of the snow algal species Chloromonas miwae (Fukushima) Muramoto et al., comb. nov. (Volvocales, Chlorophyceae) from Japan, based on molecular phylogeny and cultured material. Eur. J. Phycol. 45: 27-37.
Nakada, T., Misawa, K. and Nozaki, H. 2008a. Molecular systematics of Volvocales (Chlorophyceae, Chlorophyta) based on exhaustive 18S rRNA phylogenetic analyses. Mol. Phylogenet. Evol. 48: 281-291.
Nakada, T. and Nozaki, H. 2009. Taxonomic study of two new genera of fusiform green flagellates, Tabris gen. nov. and Hamakko gen. nov. (Volvocales, Chlorophyceae). J. Phycol. 45: 482-492.
Nakada, T., Nozaki, H. and Proschold, T. 2008b. Molecular phylogeny, ultrastructure, and taxonomic revision of Chlorogonium (Chlorophyta): Emendation of Chlorogonium and description of Gungnir gen. nov. and Rusalka gen. nov. J. Phycol. 44: 751-760.
Nakada, T., Nozaki, H. and Tomita, M. 2010. Another origin of coloniality in volvocaleans: the phylogenetic position of Pyrobotrys arnoldi (Spondylomoraceae, Volvocales). J. Eukaryot. Microbiol. 57: 379-382.
Nakayama, T., Kawachi, M. and Inouye, I. 2000. Taxonomy and the phylogenetic position of a new prasinophycean alga, Crustomastix didyma gen. & sp. nov. (Chlorophyta). Phycologia 39: 337-348.
Nakayama, T., Marin, B., Kranz, H. D., Surek, B., Huss, V. A. R., Inouye, I. and Melkonian, M. 1998. The basal position of scaly green flagellates among the green algae (Chlorophyta) is revealed by analyses of nuclear-encoded SSU rRNA sequences. Protist 149: 367-380.
Nakayama, T., Suda, S., Kawachi, M. and Inouye, I. 2007. Phylogeny and ultrastructure of Nephroselmis and Pseudoscourfieldia (Chlorophyta), including the description of Nephroselmis anterostigmatica sp. nov. and a proposal for the Nephroselmidales ord. nov. Phycologia 46: 680-697.
Nakayama, T., Watanabe, S. and Inouye, I. 1996a. Phylogeny of wall-less green flagellates inferred from 18S rDNA sequence data. Phycol. Res. 44: 151-161.
Nakayama, T., Watanabe, S., Mitsui, K., Uchida, H. and Inouye, I. 1996b. The phylogenetic relationship between the Chlamydomonadales and Chlorococcales inferred from 18S rDNA sequence data. Phycol. Res. 44: 47-55.
Nakazawa, A. S., Yamada, T. and Nozaki, H. 2004. Taxonomic study of Asterococcus (Chlorophyceae) based on comparative morphology and rbcL gene sequences. Phycologia 43: 711-721.
Nedelcu, A. M. 1997. Fragmented and scrambled mitochondrial ribosomal RNA coding regions among green algae: A model for their origin and evolution. Mol. Biol. Evol. 14: 506-517.
Nedelcu, A. M. 1998. Contrasting mitochondrial genome organizations and sequence affiliations among green algae: Potential factors, mechanisms, and evolutionary scenarios. J. Phycol. 34: 16-28.
Nedelcu, A. M. 2001. Complex patterns of plastid 16S rRNA gene evolution in nonphotosynthetic green algae. J. Mol. Evol. 53: 670-679.
Nedelcu, A. M. 2009. Environmentally induced responses co-opted for reproductive altruism. Biol. Lett. 5: 805-808.
Nedelcu, A. M., Borza, T. and Lee, R. W. 2006. A land plant-specific multigene family in the unicellular Mesostigma argues for its close relationship to Streptophyta. Mol. Biol. Evol. 23: 1011-1015.
Nedelcu, A. M. and Lee, R. W. 1998. Short repetitive sequences in green algal mitochondrial genomes: Potential roles in mitochondrial genome evolution. Mol. Biol. Evol. 15: 690-701.
Nedelcu, A. M., Lee, R. W., Lemieux, C., Gray, M. W. and Burger, G. 2000. The complete mitochondrial DNA sequence of Scenedesmus obliquus reflects an intermediate stage in the evolution of the green algal mitochondrial genome. Genome Res. 10: 819-831.
Nedelcu, A. M. and Michod, R. E. 2006. The evolutionary origin of an altruistic gene. Mol. Biol. Evol. 23: 1460-1464.
Nedelcu, A. M., Miles, I. H., Fagir, A. M. and Karol, K. 2008. Adaptive eukaryote-to-eukaryote lateral gene transfer: stress-related genes of algal origin in the closest unicellular relatives of animals. J. Evol. Biol. 21: 1852-1860.
Nelson, W. A. and Ryan, K. G. 1986. Palmophyllum umbracola sp. nov. (Chlorophyta) from offshore islands of northern New Zealand. Phycologia 25: 168-177.
Neustupa, J., Elias, M., Skaloud, P., Nemcova, Y. and Sejnohova, L. 2011. Xylochloris irregularis, gen. et sp. nov. (Trebouxiophyceae, Chlorophyta), a novel subaerial coccoid green alga. Phycologia 50: 57-66.
Neustupa, J., Nemcova, Y., Elias, M. and Skaloud, P. 2009. Kalinella bambusicola gen. et sp. nov. (Trebouxiophyceae, Chlorophyta), a novel coccoid Chlorella-like subaerial alga from Southeast Asia. Phycol. Res. 57: 159-169.
Niklas, K. J. and Kutschera, U. 2010. The evolution of the land plant life cycle. New Phytol. 185: 27-41.
Nishii, I. and Miller, S. M. 2010. Volvox: Simple steps to developmental complexity? Curr. Opin. Plant Biol. 13: 646-653.
Nishii, I., Ogihara, S. and Kirk, D. L. 2003. A kinesin, InvA, plays an essential role in Volvox morphogenesis. Cell 113: 743-753.
Noble, G., Rogers, M. and Keeling, P. 2007. Complex distribution of EFL and EF-1α proteins in the green algal lineage. BMC Evol. Biol. 7: 82.
Nosek, J. and Tomáska, L. 2003. Mitochondrial genome diversity: evolution of the molecular architecture and replication strategy. Curr. Genet. 44: 73-84.
Not, F., Latasa, M., Marie, D., Cariou, T., Vaulot, D. and Simon, N. 2004. A single species, Micromonas pusilla (Prasinophyceae), dominates the eukaryotic picoplankton in the western English channel. Appl. Environ. Microbiol. 70: 4064-4072.
Novis, P. M., Lorenz, M., Broady, P. A. and Flint, E. A. 2010. Parallela Flint: its phylogenetic position in the Chlorophyceae and the polyphyly of Radiofilum Schmidle. Phycologia 49: 373-383.
Nowack, E. C. M. and Melkonian, M. 2010. Endosymbiotic associations within protists. Philos. Trans. R. Soc. Lond. B Biol. Sci. 365: 699-712.
Nozaki, H. 1990. Ultrastructure of the extracellular matrix of Gonium (Volvocales, Chlorophyta). Phycologia 29: 1-8.
Nozaki, H. and Kuroiwa, T. 1992. Ultrastructure of the extracellular matrix and taxonomy of Eudorina, Pleodorina and Yamagishiella gen. nov. (Volvocaceae, Chlorophyta). Phycologia 31: 529-541.
Nozaki, H., Misawa, K., Kajita, T., Kato, M., Nohara, S. and Watanabe, M. M. 2000. Origin and evolution of the colonial Volvocales (Chlorophyceae) as inferred from multiple, chloroplast gene sequences. Mol. Phylogenet. Evol. 17: 256-268.
Nozaki, H., Misumi, O. and Kuroiwa, T. 2003. Phylogeny of the quadriflagellate Volvocales (Chlorophyceae) based on chloroplast multigene sequences. Mol. Phylogenet. Evol. 29: 58-66.
Nozaki, H., Mori, T., Misumi, O., Matsunaga, S. and Kuroiwa, T. 2006a. Males evolved from the dominant isogametic mating type. Curr. Biol. 16: R1018-R1020.
Nozaki, H., Nakada, T. and Watanabe, S. 2010. Evolutionary origin of Gloeomonas (Volvocales, Chlorophyceae), based on ultrastructure of chloroplasts and molecular phylogeny. J. Phycol. 46: 195-200.
Nozaki, H., Ott, F. D. and Coleman, A. W. 2006b. Morphology, molecular phylogeny and taxonomy of two new species of Pleodorina (Volvoceae, Chlorophyceae). J. Phycol. 42: 1072-1080.
O’Kelly, C. J. 1988. Division of Palmoclathrus stipitatus (Chlorophyta) vegetative cells. Phycologia 27: 248-253.
O’Kelly, C. J. 2007. The origin and early evolution of green plants. In: Evolution of primary producers in the sea. pp. 287-309. Falkowski, P. G. & Knoll, A. H. Eds., Elsevier Academic Press, Burlington, MA.
O’Kelly, C. J., Bellows, W. K. and Wysor, B. 2004a. Phylogenetic position of Bolbocoleon piliferum (Ulvophyceae, Chlorophyta): Evidence from reproduction, zoospore and gamete ultrastructure, and small subunit rRNA gene sequences. J. Phycol. 40: 209-222.
O’Kelly, C. J. and Floyd, G. L. 1984a. Correlations among patterns of sporangial structure and development, life histories, and ultrastructural features in the Ulvophyceae. In: Systematics of the green algae. pp. 121-156. Irvine, D. & John, D. Eds., Academic Press, London and Orlando.
O’Kelly, C. J. and Floyd, G. L. 1984b. Flagellar apparatus absolute orientations and the phylogeny of the green algae. Biosystems 16: 227-251.
O’Kelly, C. J., Kurihara, A., Shipley, T. C. and Sherwood, A. R. 2010. Molecular assessment of Ulva spp. (Ulvophyceae, Chlorophyta) in the Hawaiian islands. J. Phycol. 46: 728-735.
O’Kelly, C. J., Sieracki, M. E., Thier, E. C. and Hobson, I. C. 2003. A transient bloom of Ostreococcus (Chlorophyta, Prasinophyceae) in West Neck Bay, Long Island, New York. J. Phycol. 39: 850-854.
O’Kelly, C. J., Watanabe, S. and Floyd, G. L. 1994. Ultrastructure and phylogenetic relationships of Chaetopeltidales ord. nov. (Chlorophyta, Chlorophyceae). J. Phycol. 30: 118-128.
O’Kelly, C. J., Wysor, B. and Bellows, W. K. 2004b. Collinsiella (Ulvophyceae, Chlorophyta) and other ulotrichalean taxa with shell-boring sporophytes form a monophyletic clade. Phycologia 43: 41-49.
O’Kelly, C. J., Wysor, B. and Bellows, W. K. 2004c. Gene sequence diversity and the phylogenetic position of algae assigned to the genera Phaeophila and Ochlochaete (Ulvophyceae, Chlorophyta). J. Phycol. 40: 789-799.
Ohno, S. 1970. Evolution by Gene Duplication. Springer-Verlag, New York.
Okamoto, N. and Inouye, I. 2005. A secondary symbiosis in progress? Science 310: 287.
Okamoto, N. and Inouye, I. 2006. Hatena arenicola gen. et sp. nov., a Katablepharid undergoing probable plastid acquisition. Protist 157: 401-419.
Okuda, K., Mine, I., Morinaga, T. and Kuwaki, N. 1997a. Cytomorphogenesis in coenocytic green algae. V. Segregative cell division and cortical microtubules in Dictyosphaeria cavernosa (Siphonocladales, Chlorophyceae). Phycol. Res. 45: 189-196.
Okuda, K., Mine, I. and Ueno, S. 1997b. Cytomorphogenesis in coenocytic green algae. IV. The construction of cortical microtubules during lenticular cell formation in Valonia utricularis. Mem. Fac. Sci. Kochi Univ. Ser. D (Biol.) 18: 17-25.
Olovnikov, A. M. 1971. Principle of marginotomy in template synthesis of polynucleotides. Dok. Akad. Nauk SSSR 201: 1496-&.
Olsen, J. L., Stam, W. T., Berger, S. and Menzel, D. 1994. 18S rDNA and evolution in the Dasycladales (Chlorophyta): modern living fossils. J. Phycol. 30: 729-744.
Oren, A. 2005. A hundred years of Dunaliella research: 1905-2005. Saline Systems 1: 2.
Palenik, B., Grimwood, J., Aerts, A., Rouze, P., Salamov, A., et al. 2007. The tiny eukaryote Ostreococcus provides genomic insights into the paradox of plankton speciation. Proc. Natl. Acad. Sci. U.S.A. 104: 7705-7710.
Palmer, J. D. 1985. Comparative organization of chloroplast genomes. Annu. Rev. Genet. 19: 325-354.
Parke, M., Boalch, G. T., Jowett, R. and Harbour, D. S. 1978. The genus Pterosperma (Prasinophyceae): species with a single equatorial ala. J. Mar. Biol. Assoc. U. K. 58: 239-276.
Parke, M. and Manton, I. 1967. The specific identity of the algal symbiont in Convoluta roscoffensis. J. Mar. Biol. Assoc. U. K. 47: 445-464.
Pascher, A. 1914. Über Flagellaten und Algen. Ber. Dtsch. Bot. Ges. 32: 136-160.
Pazoutova, M., Skaloud, P. and Nemjova, K. 2010. Phylogenetic position of Ooplanctella planoconvexa, gen. et comb. nova and Echinocoleum elegans (Oocystaceae, Trebouxiophyceae, Chlorophyta). Fottea 10: 75-82.
Peers, G., Truong, T. B., Ostendorf, E., Busch, A., Elrad, D., Grossman, A. R., Hippler, M. and Niyogi, K. K. 2009. An ancient light-harvesting protein is critical for the regulation of algal photosynthesis. Nature 462: 518-U215.
Perasso, R., Baroin, A., Qu, L. H., Bachellerie, J. P. and Adoutte, A. 1989. Origin of the algae. Nature 339: 142-144.
Petersen, J., Teich, R., Becker, B., Cerff, R. and Brinkmann, H. 2006. The GapA/B gene duplication marks the origin of Streptophyta (charophytes and land plants). Mol. Biol. Evol. 23: 1109-1118.
Philippe, H. and Telford, M. J. 2006. Large-scale sequencing and the new animal phylogeny. Trends Ecol. Evol. 21: 614-620.
Picket-Heaps, J. D. and Marchant, H. J. 1972. The phylogeny of the green algae: A new proposal. Cytobios 6: 255-264.
Pickett-Heaps, J. D. 1975. Green algae: Structure, reproduction and evolution in selected genera. Sinauer Associates, Sunderland, Massachusetts.
Piganeau, G. and Moreau, H. 2007. Screening the Sargasso Sea metagenome for data to investigate genome evolution in Ostreococcus (Prasinophyceae, Chlorophyta). Gene 406: 184-190.
Pombert, J.-F. and Keeling, P. J. 2010. The mitochondrial genome of the entomoparasitic green alga Helicosporidium. PLoS One 5: e8954.
Pombert, J. F., Lemieux, C. and Turmel, M. 2006. The complete chloroplast DNA sequence of the green alga Oltmannsiellopsis viridis reveals a distinctive quadripartite architecture in the chloroplast genome of early diverging ulvophytes. BMC Biology 4: 3.
Pombert, J. F., Otis, C., Lemieux, C. and Turmel, M. 2004. The complete mitochondrial DNA sequence of the green alga Pseudendoclonium akinetum (Ulvophyceae) highlights distinctive evolutionary trends in the Chlorophyta and suggests a sister-group relationship between the Ulvophyceae and Chlorophyceae. Mol. Biol. Evol. 21: 922-935.
Pombert, J. F., Otis, C., Lemieux, C. and Turmel, M. 2005. The chloroplast genome sequence of the green alga Pseudendoclonium akinetum (Ulvophyceae) reveals unusual structural features and new insights into the branching order of chlorophyte lineages. Mol. Biol. Evol. 22: 1903-1918.
Popescu, C. E. and Lee, R. W. 2007. Mitochondrial genome sequence evolution in Chlamydomonas. Genetics 175: 819-826.
Popper, Z. A., Michel, G., Hervé, C., Domozych, D. S., Willats, W. G. T., Tuohy, M. G., Kloareg, B. and Stengel, D. B. 2011. Evolution and diversity of plant cell walls: From algae to flowering plants. Annu. Rev. Plant Biol. 62: 567-590.
Popper, Z. A. and Tuohy, M. G. 2010. Beyond the green: Understanding the evolutionary puzzle of plant and algal cell walls. Plant Physiol. 153: 373-383.
Prochnik, S. E., Umen, J., Nedelcu, A. M., Hallmann, A., Miller, S. M., et al. 2010. Genomic analysis of organismal complexity in the multicellular green alga Volvox carteri. Science 329: 223-226.
Proost, S., Van Bel, M., Sterck, L., Billiau, K., Van Parys, T., Van de Peer, Y. and Vandepoele, K. 2009. PLAZA: A Comparative Genomics Resource to Study Gene and Genome Evolution in Plants. Plant Cell 21: 3718-3731.
Pröschold, T., Bock, C., Luo, W. and Krienitz, L. 2010. Polyphyletic distribution of bristle formation in Chlorellaceae: Micractinium, Diacanthos, Didymogenes and Hegewaldia gen. nov. (Trebouxiophyceae, Chlorophyta). Phycol. Res. 58: 1-8.
Pröschold, T., Darienko, T., Silva, P. C., Reisser, W. and Krienitz, L. 2011. The systematics of Zoochlorella revisited employing an integrative approach. Environ. Microbiol. 13: 350-364.
Pröschold, T. and Leliaert, F. 2007. Systematics of the green algae: conflict of classic and modern approaches. In: Unravelling the algae: the past, present, and future of algal systematics. pp. 123-153. Brodie, J. & Lewis, J. Eds., Taylor and Francis.
Pröschold, T., Marin, B., Schlosser, U. G. and Melkonian, M. 2001. Molecular phylogeny and taxonomic revision of Chlamydomonas (Chlorophyta). I. Emendation of Chlamydomonas Ehrenberg and Chloromonas Gobi, and description of Oogamochlamys gen. nov. and Lobochlamys gen. nov. Protist 152: 265-300.
Pröschold, T., Surek, B., Marin, B. and Melkonian, M. 2002. Protist origin of the Ulvophyceae (Chlorophyta) revealed by SSU rDNA analyses of marine coccoid green algae. J. Phycol. 38 suppl.: 30-31.
Pueschel, C., Sullivan, K. and Ballantine, D. 1997. Ultrastructure of Verdigellas peltata (Palmellaceae, Chlorophyta), a deep-water, palmelloid alga with ferritin and trilaminar sheaths. Phycologia 36: 492-499.
Qiu, Y. L., Li, L. B., Wang, B., Chen, Z. D., Knoop, V., et al. 2006. The deepest divergences in land plants inferred from phylogenomic evidence. Proc. Natl. Acad. Sci. U.S.A. 103: 15511-15516.
Qiu, Y. L. and Palmer, J. D. 1999. Phylogeny of early land plants: insights from genes and genomes. Trends Plant Sci. 4: 26-30.
Raven, J. A. 1997. Multiple origins of plasmodesmata. Eur. J. Phycol. 32: 95-101.
Raven, J. A. 2000. Land plant biochemistry. Philos. Trans. R. Soc. Lond. B Biol. Sci. 355: 833-846.
Remias, D., Karsten, U., Lutz, C. and Leya, T. 2010. Physiological and morphological processes in the Alpine snow alga Chloromonas nivalis (Chlorophyceae) during cyst formation. Protoplasma 243: 73-86.
Rensing, S. A., Lang, D., Zimmer, A. D., Terry, A., Salamov, A., et al. 2008. The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 319: 64-69.
Rindi, F. 2011. Terrestrial green algae: systematics, biogeography and expected responses to climate change. In: Climate Change, Ecology and Systematics. pp. 201-227. Hodkinson, T., Jones, S., Waldren, S. & Parnell, J. Eds., Cambridge University Press, Cambridge.
Rindi, F., Guiry, M. D. and López-Bautista, J. M. 2008. Distribution, morphology, and phylogeny of Klebsormidium (Klebsormidiales, Charophyceae) in urban environments in Europe. J. Phycol. 44: 1529-1540.
Rindi, F., Lam, D. W. and López-Bautista, J. M. 2009. Phylogenetic relationships and species circumscription in Trentepohlia and Printzina (Trentepohliales, Chlorophyta). Mol. Phylogenet. Evol. 52: 329-339.
Rindi, F., López-Bautista, J. M., Sherwood, A. R. and Guiry, M. D. 2006. Morphology and phylogenetic position of Spongiochrysis hawaiiensis gen. et sp. nov., the first known terrestrial member of the order Cladophorales (Ulvophyceae, Chlorophyta). Int. J. Syst. Evol. Microbiol. 56: 913-922.
Rindi, F., McIvor, L., Sherwood, A. R., Friedl, T., Guiry, M. D. and Sheath, R. G. 2007. Molecular phylogeny of the green algal order Prasiolales (Trebouxiophyceae, Chlorophyta). J. Phycol. 43: 811-822.
Rindi, F., Mikhailyuk, T. I., Sluiman, H. J., Friedl, T. and López-Bautista, J. M. 2011. Phylogenetic relationships in Interfilum and Klebsormidium (Klebsormidiophyceae, Streptophyta). Mol. Phylogenet. Evol. 58: 218-231.
Rindi, F., Sherwood, A. R., McIvor, L., Friedl, T., Guiry, M. D. and Sheath, R. G. 2005. Genus and species-level phylogeny in the Prasiolales (Trebouxiophyceae, Chlorophyta). Phycologia 44: 205.
Robbens, S., Derelle, E., Ferraz, C., Wuyts, J., Moreau, H. and Van de Peer, Y. 2007a. The complete chloroplast and mitochondrial DNA sequence of Ostreococcus tauri: organelle genomes of the smallest eukaryote are examples of compaction. Mol. Biol. Evol. 24: 956-968.
Robbens, S., Petersen, J., Brinkmann, H., Rouzé, P. and Van de Peer, Y. 2007b. Unique regulation of the calvin cycle in the ultrasmall green alga Ostreococcus. J. Mol. Evol. 64: 601-604.
Roberts, A. W. and Roberts, E. 2004. Cellulose synthase (CesA) genes in algae and seedless plants. Cellulose 11: 419-435.
Rochaix, J. D. 1995. Chlamydomonas reinhardtii as the photosynthetic yeast. Annu. Rev. Genet. 29: 209-230.
Rodríguez-Ezpeleta, N., Brinkmann, H., Burey, S. C., Roure, B., Burger, G., Loffelhardt, W., Bohnert, H. J., Philippe, H. and Lang, B. F. 2005. Monophyly of primary photosynthetic eukaryotes: Green plants, red algae, and glaucophytes. Curr. Biol. 15: 1325-1330.
Rodríguez-Ezpeleta, N., Philippe, H., Brinkmann, H., Becker, B. and Melkonian, M. 2007. Phylogenetic analyses of nuclear, mitochondrial, and plastid multigene data sets support the placement of Mesostigma in the Streptophyta. Mol. Biol. Evol. 24: 723-731.
Rodriguez, F., Derelle, E., Guillou, L., Le Gall, F., Vaulot, D. and Moreau, H. 2005. Ecotype diversity in the marine picoeukaryote Ostreococcus (Chlorophyta, Prasinophyceae). Environ. Microbiol. 7: 853-859.
Rodriguez, F., Feist, S. W., Guillou, L., Harkestad, L. S., Bateman, K., Renault, T. and Mortensen, S. 2008. Phylogenetic and morphological characterisation of the green algae infesting blue mussel Mytilus edulis in the North and South Atlantic oceans. Dis. Aquat. Org. 81: 231-240.
Roger, A. J. and Hug, L. A. 2006. The origin and diversification of eukaryotes: problems with molecular phylogenetics and molecular clock estimation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 361: 1039-1054.
Rogers, C. E., Domozych, D. S., Stewart, K. D. and Mattox, K. R. 1981. The flagellar apparatus of Mesostigma viride (Prasinophyceae): multilayered structures in a scaly green flagellate. Plant Syst. Evol. 138: 247-258.
Rogers, C. E., Mattox, K. R. and Stewart, K. D. 1980. The zoospore of Chlorokybus atmophyticus, a charophyte with sarcinoid growth habit. Am. J. Bot. 67: 774-783.
Rogers, M. B., Gilson, P. R., Su, V., McFadden, G. I. and Keeling, P. J. 2007. The complete chloroplast genome of the chlorarachniophyte Bigelowiella natans: evidence for independent origins of chlorarachniophyte and euglenid secondary endosymbionts. Mol. Biol. Evol. 24: 54-62.
Romanel, E. A. C., Schrago, C. G., Counago, R. M., Russo, C. A. M. and Alves-Ferreira, M. 2009. Evolution of the B3 DNA binding superfamily: New insights into REM family gene diversification. PLoS One 4.
Round, F. E. 1984. The systematics of the Chlorophyta: an historical review leading to some modern concepts [the taxonomy of the Chlorophyta III]. In: Systematics of the green algae. pp. 1–28. Irvine, D. E. G. & John, D. M. Eds., Academic Press, London.
Rumpf, R., Vernon, D., Schreiber, D. and Birky, C. W. 1996. Evolutionary consequences of the loss of photosynthesis in Chlamydomonadaceae: Phylogenetic analysis of Rrn18 (18S rDNA) in 13 Polytoma strains (Chlorophyta). J. Phycol. 32: 119-126.
Rumpho, M. E., Worful, J. M., Lee, J., Kannan, K., Tyler, M. S., Bhattacharya, D., Moustafa, A. and Manhart, J. R. 2008. Horizontal gene transfer of the algal nuclear gene psbO to the photosynthetic sea slug Elysia chlorotica. Proc. Natl. Acad. Sci. U.S.A. 105: 17867-17871.
Sager, R. and Granick, S. 1954. Nutritional control of sexuality in Chlamydomonas reinhardi. J. Gen. Physiol. 37: 729-742.
Sakayama, H. 2008. Taxonomy of Nitella (Charales, Charophyceae) based on comparative morphology of oospores and multiple DNA marker phylogeny using cultured material. Phycol. Res. 56: 202-215.
Sakayama, H., Hara, Y., Arai, S., Sato, H. and Nozaki, H. 2004. Phylogenetic analyses of Nitella subgenus Tieffallenia (Charales, Charophyceae) using nuclear ribosomal DNA internal transcribed spacer sequences. Phycologia 43: 672-681.
Sakayama, H., Kasai, F., Nozaki, H., Watanabe, M. M., Kawachi, M., Shigyo, M., Nishihiro, J., Washitani, I., Krienitz, L. and Ito, M. 2009. Taxonomic reexamination of Chara globularis (Charales, Charophyceae) from Japan based on oospore morphology and rbcL gene sequences, and the description of C. leptospora sp. nov. J. Phycol. 45: 917-927.
Sakayama, H., Miyaji, K., Nagumo, T., Kato, M., Hara, Y. and Nozaki, H. 2005. Taxonomic reexamination of 17 species of Nitella subgenus Tieffallenia (Charales, Charophyceae) based on internal morphology of the oospore wall and multiple DNA marker sequences. J. Phycol. 41: 195-211.
Sanchez-Puerta, M. V., Leonardi, P. I., O’Kelly, C. J. and Cáceres, E. J. 2006. Pseudulvella americana belongs to the order Chaetopeltidales (class Chlorophyceae), evidence from ultrastructure and SSU rDNA sequence data. J. Phycol. 42: 943-950.
Santos, M. A. S., Moura, G., Massey, S. E. and Tuite, M. F. 2004. Driving change: the evolution of alternative genetic codes. Trends Genet. 20: 95-102.
Schaack, S., Gilbert, C. and Feschotte, C. 2010. Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution. Trends Ecol. Evol. 25: 537-546.
Schmidt, S. K., Lynch, R. C., King, A. J., Karki, D., Robeson, M. S., Nagy, L., Williams, M. W., Mitter, M. S. and Freeman, K. R. 2011. Phylogeography of microbial phototrophs in the dry valleys of the high Himalayas and Antarctica. Proceedings of the Royal Society B: Biological Sciences.
Schneider, S. U. and Degroot, E. J. 1991. Sequences of two rbcS cDNA clones of Batophora oerstedii: structural and evolutionary considerations. Curr. Genet. 20: 173-175.
Schneider, S. U., Leible, M. B. and Yang, X. P. 1989. Strong homology between the small subunit of ribulose-1,5-bisphosphate carboxylase oxygenase of 2 species of Acetabularia and the occurrence of unusual codon usage. Mol. Gen. Genet. 218: 445-452.
Schultz, D. W. and Yarus, M. 1994. Transfer RNA mutation and the malleability of the genetic code. J. Mol. Biol. 235: 1377-1380.
Schwartz, J. A., Curtis, N. E. and Pierce, S. K. 2010. Using algal transcriptome sequences to identify transferred genes in the sea slug, Elysia chlorotica. Evol. Biol. 37: 29-37.
Shendure, J. and Ji, H. 2008. Next-generation DNA sequencing. Nat. Biotechnol. 26: 1135-1145.
Shepherd, V. A., Beilby, M. J. and Bisson, M. A. 2004. When is a cell not a cell? A theory relating coenocytic structure to the unusual electrophysiology of Ventricaria ventricosa (Valonia ventricosa). Protoplasma 223: 79-91.
Shi, X. L., Marie, D., Jardillier, L., Scanlan, D. J. and Vaulot, D. 2009. Groups without cultured representatives dominate eukaryotic picophytoplankton in the oligotrophic South East Pacific Ocean. PLoS One 4: e7657.
Shimada, S., Yokoyama, N., Arai, S. and Hiraoka, M. 2008. Phylogeography of the genus Ulva (Ulvophyceae, Chlorophyta), with special reference to the Japanese freshwater and brackish taxa. J. Appl. Phycol. 20: 979-989.
Shoup, S. and Lewis, L. A. 2003. Polyphyletic origin of parallel basal bodies in swimming cells of Chlorophycean green algae (Chlorophyta). J. Phycol. 39: 789-796.
Simon, A., Glockner, G., Felder, M., Melkonian, M. and Becker, B. 2006. EST analysis of the scaly green flagellate Mesostigma viride (Streptophyta): Implications for the evolution of green plants (Viridiplantae). BMC Plant Biol. 6: 2.
Simon, L., Bousquet, J., Levesque, R. C. and Lalonde, M. 1993. Origin and diversification of endomycorrhizal fungi and coincidence with vascular land plants. Nature 363: 67-69.
Simpson, C. L. and Stern, D. B. 2002. The treasure trove of algal chloroplast genomes. Surprises in architecture and gene content, and their functional implications. Plant Physiol. 129: 957-966.
Six, C., Sherrard, R., Lionard, M., Roy, S. and Campbell, D. A. 2009. Photosystem II and pigment dynamics among ecotypes of the green alga Ostreococcus. Plant Physiol. 151: 379-390.
Six, C., Worden, A. Z., Rodriguez, F., Moreau, H. and Partensky, F. 2005. New insights into the nature and phylogeny of prasinophyte antenna proteins: Ostreococcus tauri, a case study. Mol. Biol. Evol. 22: 2217-2230.
Škaloud, P. and Peksa, O. 2010. Evolutionary inferences based on ITS rDNA and actin sequences reveal extensive diversity of the common lichen alga Asterochloris (Trebouxiophyceae, Chlorophyta). Mol. Phylogenet. Evol. 54: 36-46.
Škaloud, P., Kalina, T., Nemjová, K., De Clerck, O. and Leliaert, F. 2013. Morphology and phylogenetic position of the freshwater green microalgae Chlorochytrium (Chlorophyceae) and Scotinosphaera (Scotinosphaerales, ord. nov., Ulvophyceae). J. Phycol. 49: 115-129.
Šlapeta, J., Lopez-Garcia, P. and Moreira, D. 2006. Global dispersal and ancient cryptic species in the smallest marine eukaryotes. Mol. Biol. Evol. 23: 23-29.
Sleigh, M. A. 1989. Protozoa and Other Protists. Edward Arnold Limited, New York.
Sluiman, H. J. 1989. The green algal class Ulvophyceae. An ultrastructural survey and classification. Crypt. Bot. 1: 83-94.
Sluiman, H. J., Guihal, C. and Mudimu, O. 2008. Assessing phylogenetic affinities and species delimitations in Klebsormidiales (Streptophyta): Nuclear-encoded rDNA phylogenies and its secondary structure models in Klebsormidium, Hormidiella, and Entransia. J. Phycol. 44: 183-195.
Sluiman, H. J., Roberts, K. R., Stewart, K. D. and Mattox, K. R. 1983. Comparative cytology and taxonomy of the Ulvophyceae. IV. Mitosis and cytokinesis in Ulothrix. Acta Bot. Neerl. 32: 257-269.
Smith, D. R. 2009. Unparalleled GC content in the plastid DNA of Selaginella. Plant Mol. Biol. 71: 627-639.
Smith, D. R., Hua, J. M. and Lee, R. W. 2010a. Evolution of linear mitochondrial DNA in three known lineages of Polytomella. Curr. Genet. 56: 427-438.
Smith, D. R. and Lee, R. W. 2008. Mitochondrial genome of the colorless green alga Polytomella capuana: A linear molecule with an unprecedented GC content. Mol. Biol. Evol. 25: 487-496.
Smith, D. R. and Lee, R. W. 2009. The mitochondrial and plastid genomes of Volvox carteri: bloated molecules rich in repetitive DNA. BMC Genomics 10: 132.
Smith, D. R. and Lee, R. W. 2010. Low nucleotide diversity for the expanded organelle and nuclear genomes of Volvox carteri supports the mutational-hazard hypothesis. Mol. Biol. Evol. 27: 2244-2256.
Smith, D. R., Lee, R. W., Cushman, J. C., Magnuson, J. K., Tran, D. and Polle, J. E. W. 2010b. The Dunaliella salina organelle genomes: large sequences, inflated with intronic and intergenic DNA. BMC Plant Biol. 10: 83.
Soltis, D. E., Moore, M. J., Burleigh, G. and Soltis, P. S. 2009. Molecular markers and concepts of plant evolutionary relationships: Progress, promise, and future prospects. Crit. Rev. Plant Sci. 28: 1-15.
Sorensen, I., Domozych, D. and Willats, W. G. T. 2010. How have plant cell walls evolved? Plant Physiol. 153: 366-372.
Starr, R. 1969. Structure, reproduction and differentiation in Volvox carteri f. nagariensis Iyengar, strains HK9 and HK10. Arch. Protistenk. 111: 204-222.
Steemans, P., Le Herisse, A., Melvin, J., Miller, M. A., Paris, F., Verniers, J. and Wellman, C. H. 2009. Origin and radiation of the earliest vascular land plants. Science 324: 353-353.
Stein, J. R. 1965. On cytoplasmic strands in Gonium pectorale (Volvocales). J. Phycol. 1: 1-5.
Steinkötter, J., Bhattacharya, D., Semmelroth, I., Bibeau, C. and Melkonian, M. 1994. Prasinophytes form independent lineages within the Chlorophyta: evidence from ribosomal RNA sequence comparison. J. Phycol. 30: 340-345.
Stiller, J. W., Huang, J. L., Ding, Q., Tian, J. and Goodwillie, C. 2009. Are algal genes in nonphotosynthetic protists evidence of historical plastid endosymbioses? BMC Genomics 10: 484.
Suda, S., Watanabe, M. M. and Inouye, I. 2004. Electron microscopy of sexual reproduction in Nephroselmis olivacea (Prasinophyceae, Chlorophyta). Phycol. Res. 52: 273-283.
Sudman, M. S. 1974. Protothecosis: Critical review. Am. J. Clin. Pathol. 61: 10-19.
Summerer, M., Sonntag, B. and Sommaruga, R. 2008. Ciliate-symbiont specificity of freshwater endosymbiotic Chlorella (Trebouxiophyceae, Chlorophyta). J. Phycol. 44: 77-84.
Sun, G. L., Yang, Z. F., Ishwar, A. and Huang, J. L. 2010. Algal genes in the closest relatives of animals. Mol. Biol. Evol. 27: 2879-2889.
Suutari, M., Majaneva, M., Fewer, D., Voirin, B., Aiello, A., Friedl, T., Chiarello, A. and Blomster, J. 2010. Molecular evidence for a diverse green algal community growing in the hair of sloths and a specific association with Trichophilus welckeri (Chlorophyta, Ulvophyceae). BMC Evol. Biol. 10: 86.
Sweeney, B. M. 1976. Pedinomonas noctilucae (Prasinophyceae), the flagellate symbiont in Noctiluca (Dinophyceae) in Southeast Asia. J. Phycol. 12: 460-464.
Sym, S. D. and Pienaar, R. N. 1993. The class Prasinophyceae. In: Progress in phycological research. pp. 281-376. Round, F. E. & Chapman, D. J. Eds., Biopress Ltd., Bristol.
Tafresh, A. H. and Shariati, M. 2009. Dunaliella biotechnology: methods and applications. J. Appl. Microbiol. 107: 14-35.
Takahashi, F., Okabe, Y., Nakada, T., Sekimoto, H., Ito, M., Kataoka, H. and Nozaki, H. 2007. Origins of the secondary plastids of Euglenophyta and Chlorarachniophyta as revealed by an analysis of the plastid-targeting, nuclear-encoded gene psbO. J. Phycol. 43: 1302-1309.
Takishita, K., Kawachi, M., Noel, M. H., Matsumoto, T., Kakizoe, N., Watanabe, M. M., Inouye, I., Ishida, K. I., Hashimoto, T. and Inagaki, Y. 2008. Origins of plastids and glyceraldehyde-3-phosphate dehydrogenase genes in the green-colored dinoflagellate Lepidodinium chlorophorum. Gene 410: 26-36.
Tam, L. W. and Kirk, D. L. 1991. Identification of cell-type-specific genes of Volvox carteri and characterization of their expression during the asexual life cycle. Dev. Biol. 145: 51-66.
Tanabe, Y., Hasebe, M., Sekimoto, H., Nishiyama, T., Kitani, M., Henschel, K., Munster, T., Theissen, G., Nozaki, H. and Ito, M. 2005. Characterization of MADS-box genes in charophycean green algae and its implication for the evolution of MADS-box genes. Proc. Natl. Acad. Sci. U.S.A. 102: 2436-2441.
Tappan, H. 1980. Palaeobiology of Plant Protists. Freeman, San Francisco.
Tartar, A. and Boucias, D. G. 2004. The non-photosynthetic, pathogenic green alga Helicosporidium sp. has retained a modified, functional plastid genome. FEMS Microbiol. Lett. 233: 153-157.
Tartar, A., Boucias, D. G., Becnel, J. J. and Adams, B. J. 2003. Comparison of plastid 16S rRNA (rrn 16) genes from Helicosporidium spp.: evidence supporting the reclassification of Helicosporidia as green algae (Chlorophyta). Int. J. Syst. Evol. Microbiol. 53: 1719-1723.
Timme, R. E. and Delwiche, C. F. 2010. Uncovering the evolutionary origin of plant molecular processes: comparison of Coleochaete (Coleochaetales) and Spirogyra (Zygnematales) transcriptomes. BMC Plant Biol. 10: 96.
* Timme, R. E., Bachvaroff, T. R. and Delwiche, C. F. 2012. Broad phylogenomic sampling and the sister lineage of land plants. PLoS One 7: e29696.
Tirichine, L. and Bowler, C. 2011. Decoding algal genomes: tracing back the history of photosynthetic life on Earth. The Plant Journal 66: 45-57.
* Tippery, N. P., Fu?íková, K., Lewis, P. O. and Lewis, L. A. 2012. Probing the monophyly of the Sphaeropleales (Chlorophyceae) using data from five genes. J. Phycol. 48: 1482-1493.
Tremouillaux-Guiller, J., Rohr, T., Rohr, R. and Huss, V. A. R. 2002. Discovery of an endophytic alga in Ginkgo biloba. Am. J. Bot. 89: 727-733.
Triemer, R. and Farmer, M. 2007. A decade of euglenoid molecular phylogenetics. In: Unravelling the algae: the past, present, and future of algal systematics. pp. 315-330. Brodie, J. & Lewis, J. Eds., CRC Press, Taylor and Francis, Boca Raton.
Tsekos, I. 1999. The sites of cellulose synthesis in algae: diversity and evolution of cellulose-synthesizing enzyme complexes. J. Phycol. 35: 635-655.
Turmel, M., Brouard, J.-S., Gagnon, C., Otis, C. and Lemieux, C. 2008. Deep division in the Chlorophyceae (Chlorophyta) revealed by chloroplast phylogenomic analyses. J. Phycol. 44: 739-750.
Turmel, M., Ehara, M., Otis, C. and Lemieux, C. 2002a. Phylogenetic relationships among streptophytes as inferred from chloroplast small and large subunit rRNA gene sequences. J. Phycol. 38: 364-375.
Turmel, M., Gagnon, M.-C., O’Kelly, C. J., Otis, C. and Lemieux, C. 2009a. The chloroplast genomes of the green algae Pyramimonas, Monomastix, and Pycnococcus shed new light on the evolutionary history of prasinophytes and the origin of the secondary chloroplasts of euglenids. Mol. Biol. Evol. 26: 631-648.
Turmel, M., Lemieux, C., Burger, G., Lang, B. F., Otis, C., Plante, I. and Gray, M. W. 1999a. The complete mitochondrial DNA sequences of Nephroselmis olivacea and Pedinomonas minor: Two radically different evolutionary patterns within green algae. Plant Cell 11: 1717-1729.
Turmel, M., Otis, C., Lemieux and C. 1999b. The complete chloroplast DNA sequence of the green alga Nephroselmis olivacea: Insights into the architecture of ancestral chloroplast genomes. Proc. Natl. Acad. Sci. U.S.A. 96: 10248-10253.
Turmel, M., Otis, C. and Lemieux, C. 2002b. The chloroplast and mitochondrial genome sequences of the charophyte Chaetosphaeridium globosum: Insights into the timing of the events that restructured organelle DNAs within the green algal lineage that led to land plants. Proc. Natl. Acad. Sci. U.S.A. 99: 11275-11280.
Turmel, M., Otis, C. and Lemieux, C. 2002c. The complete mitochondrial DNA sequence of Mesostigma viride identifies this green alga as the earliest green plant divergence and predicts a highly compact mitochondrial genome in the ancestor of all green plants. Mol. Biol. Evol. 19: 24-38.
Turmel, M., Otis, C. and Lemieux, C. 2003. The mitochondrial genome of Chara vulgaris: Insights into the mitochondrial DNA architecture of the last common ancestor of green algae and land plants. Plant Cell 15: 1888-1903.
Turmel, M., Otis, C. and Lemieux, C. 2005. The complete chloroplast DNA sequences of the charophycean green algae Staurastrum and Zygnema reveal that the chloroplast genome underwent extensive changes during the evolution of the Zygnematales. BMC Biology 3: 22.
Turmel, M., Otis, C. and Lemieux, C. 2006a. The chloroplast genome sequence of Chara vulgaris sheds new light into the closest green algal relatives of land plants. Mol. Biol. Evol. 23: 1324-1338.
Turmel, M., Otis, C. and Lemieux, C. 2006b. The chloroplast genome sequence of Chara vulgaris sheds new light into the closest green algal relatives of land plants. Mol. Biol. Evol. 23: 1324-1338.
Turmel, M., Otis, C. and Lemieux, C. 2007a. An unexpectedly large and loosely packed mitochondrial genome in the charophycean green alga Chlorokybus atmophyticus. BMC Genomics 8: 137.
Turmel, M., Otis, C. and Lemieux, C. 2009b. The chloroplast genomes of the green algae Pedinomonas minor, Parachlorella kessleri, and Oocystis solitatia reveal a shared ancestry between the Pedinomonadales and Chlorellales. Mol. Biol. Evol. 26: 2317-2331.
Turmel, M., Otis, C. and Lemieux, C. 2010. A deviant genetic code in the reduced mitochondrial genome of the picoplanktonic green alga Pycnococcus provasolii. J. Mol. Evol. 70: 203-214.
Turmel, M., Pombert, J. F., Charlebois, P., Otis, C. and Lemieux, C. 2007b. The green algal ancestry of land plants as revealed by the chloroplast genome. Int. J. Plant Sci. 168: 679-689.
Tyler, B. M., Tripathy, S., Zhang, X. M., Dehal, P., Jiang, R. H. Y., et al. 2006. Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science 313: 1261-1266.
Ueki, N., Matsunaga, S., Inouye, I. and Hallmann, A. 2010. How 5000 independent rowers coordinate their strokes in order to row into the sunlight: Phototaxis in the multicellular green alga Volvox. BMC Biology 8: 103.
Ueki, N. and Nishii, I. 2009. Controlled enlargement of the glycoprotein vesicle surrounding a Volvox embryo requires the invB nucleotide-sugar transporter and is required for normal morphogenesis. Plant Cell 21: 1166-1181.
Ueno, R., Hanagata, N., Urano, N. and Suzuki, M. 2005. Molecular phylogeny and phenotypic variation in the heterotrophic green algal genus Prototheca (Trebouxiophyceae, Chlorophyta). J. Phycol. 41: 1268-1280.
Ueno, R., Urano, N. and Suzuki, M. 2003. Phylogeny of the non-photosynthetic green micro-algal genus Prototheca (Trebouxiophyceae, Chlorophyta) and related taxa inferred from SSU and LSU ribosomal DNA partial sequence data. FEMS Microbiol. Lett. 223: 275-280.
Vahrenholz, C., Riemen, G., Pratje, E., Dujon, B. and Michaelis, G. 1993. Mitochondrial DNA of Chlamydomonas reinhardtii: the structure of the ends of the linear 15.8-kb genome suggests mechanisms for DNA replication. Curr. Genet. 24: 241-247.
van den Hoek, C., Mann, D. G. and Jahns, H. M. 1995. Algae: an introduction to phycology. Cambridge University Press.
van den Hoek, C., Stam, W. T. and Olsen, J. L. 1988. The emergence of a new chlorophytan system, and Dr. Kornmann’s contribution thereto. Helgol. Meeresunters. 42: 339-383.
Van Sandt, V. S. T., Stieperaere, H., Guisez, Y., Verbelen, J.-P. and Vissenberg, K. 2007. XET activity is found near sites of growth and cell elongation in bryophytes and some green algae: New insights into the evolution of primary cell wall elongation. Annals of Botany 99: 39-51.
Vaulot, D., Eikrem, W., Viprey, M. and Moreau, H. 2008. The diversity of small eukaryotic phytoplankton (≤ 3 µm) in marine ecosystems. FEMS Microbiol. Rev. 32: 795-820.
Verbruggen, H., Ashworth, M., LoDuca, S. T., Vlaeminck, C., Cocquyt, E., Sauvage, T., Zechman, F. W., Littler, D. S., Littler, M. M., Leliaert, F. and De Clerck, O. 2009a. A multi-locus time-calibrated phylogeny of the siphonous green algae. Mol. Phylogenet. Evol. 50: 642-653.
Verbruggen, H., Vlaeminck, C., Sauvage, T., Sherwood, A. R., Leliaert, F. and De Clerck, O. 2009b. Phylogenetic analysis of Pseudochlorodesmis strains reveals cryptic diversity above the family level in the siphonous green algae (Bryopsidales, Chlorophyta). J. Phycol. 45: 726-731.
Verghese, B. 2007. Phylogeny and evolution of the Chlorophyceae and Trebouxiophyceae. Ph.D., The University of Tulsa, 138 pp.
Vernon, D., Gutell, R. R., Cannone, J. J., Rumpf, R. W. and William Birky Jr, C. 2001. Accelerated Evolution of Functional Plastid rRNA and Elongation Factor Genes Due to Reduced Protein Synthetic Load After the Loss of Photosynthesis in the Chlorophyte Alga Polytoma. Mol. Biol. Evol. 18: 1810-1822.
Vinogradova, O. M. and Darienko, T. M. 2008. Terrestrial algae of hypersaline environments of the Central Syvash islands (Kherson Region, Ukraine). Biologia 63: 813-823.
Viprey, M., Guillou, L., Ferréol, M. and Vaulot, D. 2008. Wide genetic diversity of picoplanktonic green algae (Chloroplastida) in the Mediterranean Sea uncovered by a phylum-biased PCR approach. Environ. Microbiol. 10: 1804-1822.
Vroom, P. S. and Smith, C. M. 2001. The challenge of siphonous green algae. Am. Sci. 89: 524-531.
Wada, M., Kagawa, T. and Sato, Y. 2003. Chloroplast movement. Annu. Rev. Plant Biol. 54: 455-468.
Wägele, H., Deusch, O., Händeler, K., Martin, R., Schmitt, V., Christa, G., Pinzger, B., Gould, S. B., Dagan, T., Klussmann-Kolb, A. and Martin, W. 2011. Transcriptomic evidence that longevity of acquired plastids in the photosynthetic slugs Elysia timida and Plakobranchus ocellatus does not entail lateral transfer of algal nuclear genes. Mol. Biol. Evol. 28: 699-706.
Wakasugi, T., Nagai, T., Kapoor, M., Sugita, M., Ito, M., et al. 1997. Complete nucleotide sequence of the chloroplast genome from the green alga Chlorella vulgaris: The existence of genes possibly involved in chloroplast division. Proc. Natl. Acad. Sci. U.S.A. 94: 5967-5972.
Ward, B. L., Anderson, R. S. and Bendich, A. J. 1981. The mitochondrial genome is large and variable in a family of plants (Cucurbitaceae). Cell 25: 793-803.
Watanabe, M. M., Suda, S., Inouye, I., Sawaguchi, T. and Chihara, M. 1990. Lepidodinium viride gen. et sp. nov. (Gymnodiniales, Dinophyta), a green dinoflagellate with a chlorophyll a-containing and b-containing endosymbiont. J. Phycol. 26: 741-751.
Watanabe, S. and Floyd, G. L. 1989. Ultrastructure of the quadriflagellate zoospores of the filamentous green algae Chaetophora incrassata and Pseudoschizomeris caudata (Chaetophorales, Chlorophyceae) with emphasis on the flagellar apparatus. Bot. Mag. Tokyo 102: 533-546.
Watanabe, S., Himizu, A., Lewis, L. A., Floyd, G. L. and Fuerst, P. A. 2000. Pseudoneochloris marina (Chlorophyta), a new coccoid ulvophycean alga, and its phylogenetic position inferred from morphological and molecular data. J. Phycol. 36: 596-604.
Watanabe, S., Kuroda, N. and Maiwa, F. 2001. Phylogenetic status of Helicodictyon planctonicum and Desmochloris halophila gen. et comb. nov. and the definition of the class Ulvophyceae (Chlorophyta). Phycologia 40: 421-434.
Watanabe, S., Mitsui, K., Nakayama, T. and Inouye, I. 2006a. Phylogenetic relationships and taxonomy of sarcinoid green algae: Chlorosarcinopsis, Desmotetra, Sarcinochlamys gen. nov., Neochlorosarcina, and Chlorosphaeropsis (Chlorophyceae, Chlorophyta). J. Phycol. 42: 679-695.
Watanabe, S. and Nakayama, T. 2007. Ultrastructure and phylogenetic relationships of the unicellular green algae Ignatius tetrasporus and Pseudocharacium americanum (Chlorophyta). Phycol. Res. 55: 1-16.
Watanabe, S., Tsujimura, S., Misono, T., Nakamura, S. and Inoue, H. 2006b. Hemiflagellochloris kazakhstanica gen. et sp. nov.: A new coccoid green alga with flagella of considerably unequal lengths from a saline irrigation land in Kazakhstan (Chlorophyceae, Chlorophyta). J. Phycol. 42: 696-706.
Waters, E. R. 2003. Molecular adaptation and the origin of land plants. Mol. Phylogenet. Evol. 29: 456-463.
Watson, J. D. 1972. Origin of concatemeric T7 DNA. Nat. New Biol. 239: 197-201.
Wehr, J. D. and Sheath, R. G. 2003. Freshwater Algae of North America: Ecology and Classification. Academic Press, New York.
Wheeler, G. L. and Brownlee, C. 2008. Ca2+ signalling in plants and green algae – changing channels. Trends Plant Sci. 13: 506-514.
Wilcox, L. W., Lewis, L. A., Fuerst, P. A. and Floyd, G. L. 1992. Assessing the relationships of autosporic and zoosporic chlorococcalean green-algae with 18S rDNA sequence data. J. Phycol. 28: 381-386.
Williamson, C. E. 1979. Ultrastructural investigation of algal symbiosis in white and green Spongilla lacustris (L.) (Porifera: Spongillidae). Trans. Am. Microsc. Soc. 98: 59-77.
Winands, A. and Wagner, G. 1996. Phytochrome of the green alga Mougeotia: cDNA sequence, autoregulation and phylogenetic position. Plant Mol. Biol. 32: 589-597.
Wodniok, S., Brinkmann, H., Glockner, G., Heidel, A., Philippe, H., Melkonian, M. and Becker, B. 2011. Origin of land plants: Do conjugating green algae hold the key? BMC Evol. Biol. 11: 104.
Wodniok, S., Simon, A., Glockner, G. and Becker, B. 2007. Gain and loss of polyadenylation signals during evolution of green algae. BMC Evol. Biol. 7: 65.
Wolf, M., Buchheim, M., Hegewald, E., Krienitz, L. and Hepperle, D. 2002. Phylogenetic position of the Sphaeropleaceae (Chlorophyta). Plant Syst. Evol. 230: 161-171.
Wolf, M., Hepperle, D. and Krienitz, L. 2003. On the phylogeny of Radiococcus, Planktosphaeria and Schizochlamydella (Radiococcaceae, Chlorophyta). Biologia 58: 759-765.
Wolfe, K. H., Li, W. H. and Sharp, P. M. 1987. Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc. Natl. Acad. Sci. U.S.A. 84: 9054-9058.
Wolfe, K. H., Sharp, P. M. and Li, W. H. 1989. Rates of synonymous substitution in plant nuclear genes. J. Mol. Evol. 29: 208-211.
Wolff, G., Plante, I., Lang, B. F., Kuck, U. and Burger, G. 1994. Complete sequence of the mitochondrial DNA of the chlorophyte alga Prototheca wickerhamii. Gene content and genome organization. J. Mol. Biol. 237: 75-86.
Womersley, H. B. S. 1984. The Marine Benthic Flora of Southern Australia. Part I. Government Printer, South Australia, Adelaide.
Worden, A. Z., Lee, J. H., Mock, T., Rouze, P., Simmons, M. P., et al. 2009. Green evolution and dynamic adaptations revealed by genomes of the marine picoeukaryotes Micromonas. Science 324: 268-272.
Worden, A. Z., Nolan, J. K. and Palenik, B. 2004. Assessing the dynamics and ecology of marine picophytoplankton: The importance of the eukaryotic component. Limnol. Oceanogr. 49: 168-179.
Yamada, T. K., Miyaji, K. and Nozaki, H. 2008. A taxonomic study of Eudorina unicocca (Volvocaceae, Chlorophyceae) and related species, based on morphology and molecular phylogeny. Eur. J. Phycol. 43: 317-326.
Yin, Y. B., Huang, J. L. and Xu, Y. 2009. The cellulose synthase superfamily in fully sequenced plants and algae. BMC Plant Biol. 9: 99.
Yoon, H. S., Hackett, J. D., Ciniglia, C., Pinto, G. and Bhattacharya, D. 2004. A molecular timeline for the origin of photosynthetic eukaryotes. Mol. Biol. Evol. 21: 809-818.
Yoshii, Y., Takaichi, S., Maoka, T. and Inouye, I. 2003. Photosynthetic pigment composition in the primitive green alga Mesostigma viride (Prasinophyceae): Phylogenetic and evolutionary implications. J. Phycol. 39: 570-576.
Zechman, F. W. 2003. Phylogeny of the Dasycladales (Chlorophyta, Ulvophyceae) based on analyses of RUBISCO large subunit (rbcL) gene sequences. J. Phycol. 39: 819-827.
Zechman, F. W., Theriot, E. C., Zimmer, E. A. and Chapman, R. L. 1990. Phylogeny of the Ulvophyceae (Chlorophyta): cladistic analysis of nuclear-encoded rRNA sequence data. J. Phycol. 26: 700-710.
Zechman, F. W., Verbruggen, H., Leliaert, F., Ashworth, M., Buchheim, M. A., Fawley, M. W., Spalding, H., Pueschel, C. M., Buchheim, J. A., Verghese, B. and Hanisak, M. D. 2010. An unrecognized ancient lineage of green plants persists in deep marine waters. J. Phycol. 46: 1288-1295.
Zettler, L. A. A., Gomez, F., Zettler, E., Keenan, B. G., Amils, R. and Sogin, M. L. 2002. Eukaryotic diversity in Spain’s River of Fire. Nature 417: 137.
Zhang, J. M., Huss, V. A. R., Sun, X. P., Chang, K. J. and Pang, D. B. 2008. Morphology and phylogenetic position of a trebouxiophycean green alga (Chlorophyta) growing on the rubber tree, Hevea brasiliensis , with the description of a new genus and species. Eur. J. Phycol. 43: 185-193.
Zhu, F., Massana, R., Not, F., Marie, D. and Vaulot, D. 2005. Mapping of picoeucaryotes in marine ecosystems with quantitative PCR of the 18S rRNA gene. FEMS Microbiol. Ecol. 52: 79-92.
Ziemann, M., Bhave, M. and Zachgo, S. 2009. Origin and diversification of land plant CC-type glutaredoxins. Genome Biol. Evol. 1: 265-277.
Zingone, A., Borra, M., Brunet, C., Forlani, G., Kooistra, W. H. C. F. and Procaccini, G. 2002. Phylogenetic position of Crustomastix stigmatica sp. nov. and Dolichomastix tenuilepis in relation to the Mamiellales (Prasinophyceae, Chlorophyta). J. Phycol. 38: 1024-1039.
Zuccarello, G. C., Price, N., Verbruggen, H. and Leliaert, F. 2009. Analysis of a plastid multigene data set and the phylogenetic position of the marine macroalga Caulerpa filiformis (Chlorophyta). J. Phycol. 45: 1206-1212.
Last updated: 21 May 2013

Hello.This article was really motivating, particularly because I was investigating for thoughts on this topic last Friday.
Really good blog,thank so much for your effort in writing the posts.
Very good site,thank so much for your effort in writing the posts.
Hello to all the authors.
Very informative research.
Can I request the use of this research in my course for which I teach?
The original images in either ppt or pdf format would really be helpful. Credits of your research will always be on display when I present. It might help with future citations? – you never know.
If this request is not within the rules please let me know. However, I like to promote good research for the students.
Best
Mark